67 A/CHICKEN/VIETNAM/NCVD-04/2008
52
94
94
83
91
A/CHICKEN/VIETNAM/NCVD-05/2008 A/CHICKEN/VIETNAM/NCVD-093/2008 A/CHICKEN/VIETNAM/NCVD-03/2008
A/CHICKEN/VIETNAM/NCVD-016/2008
Clade 7
83
35
A/chicken/Shanxi/10/2006 2006
A/chicken/Shandong/A-10/2006 2006
A/chicken/Shanxi/2/2006
Clade 7
33
33
A/Beijing/01/2003
A/chicken/Korea/es/2003
99
A/black-headed goose/Qinghai/January 2005
A/chicken/Bhutan/248006/2010 A/duck/India/TR-NIV4396/2008
A/duck/Hunan/1386/2003
99
14
59
55
58
27
A/Indonesia/239H/2005 A/Indonesia/CDC1046/2007 A/Indonesia/535H/2006
A/Ck/Indonesia/2A/2003 2003 A/chicken/East Java/UT6045/2007
A/duck/Guangxi/351/2004
96
28 80 A/chicken/Thailand/ICRC-7356/2010
68
53
34
51
53
66
74
31
A/Thailand/Kan353/2004 2004
A/duck/Vietnam/48/2004 2004 A/Cambodia/P0322095/2005
A/Vietnam/CL100/2004 A/duck/Vietnam/286/2005
A/Muscovy duck/Ca Mau/April 7, 2007 A/chicken/Lang Son/200/2005 2005
A/duck/Guiyang/3834/2005 2005
A/Anhui/T2/2006 2006
46
77 A/Anhui/2/2005 2005
25
68
A/Shanghai/1/2006 2006
A/crested myna/Hong Kong/540/2006
41
99
97
33
A/Hong Kong/6841/2010 A/mallard/Korea/1195/2010
A/Hunan/1/2006
83
77 A/duck/Guangxi/150/2006
A/duck/Laos/P0117/2007
A/duck/Zhejiang/52/2000
A/goose/Fujian/bb/2003
100
A/goose/Vietnam/3/05 2005
99 A/Duck/Hong Kong/y283/1997 A/Hong Kong/156/1997
94
93
A/duck/Shantou/195/2001 A/Chicken/Hong Kong/SF219/2001
A/Goose/Guangdong/1/96
0.002
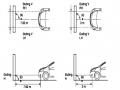
Figure 3.5. Phylogenetic tree of M gene nucleotide sequence of A/H5N1 clade 7 strains (Established by neighbor-joining analysis using MEGA 5 software, boostrap value 1000)
Table 3.7. Identification of A/H5N1 strains used in the establishment of the M-genotype
STT
International trade | Species battery | Capacity set up | Place of isolation | S bank registration gene | |
1 | A/Goose/Guangdong/1/96 | Goose | 1996 | China | AF144306 |
2 | A/Hong Kong/156/1997 | People | 1997 | Hong Kong | AF036358 |
3 | A/Duck/Hong Kong/y283/1997 | Duck | 1997 | Hong Kong | AF098567 |
4 | A/duck/Zhejiang/52/2000 | Duck | 2000 | China | AY585397 |
5 | A/duck/Shantou/195/2001 | Duck | 2001 | Hong Kong | CY029002 |
6 | A/Chicken/Hong Kong/SF219/2001 | Chicken | 2001 | China | AF509048, |
7 | A/chicken/Korea/es/2003 | Chicken | 2003 | Korea | EF541460 |
8 | A/Ck/Indonesia/2A/2003 | Chicken | 2003 | Indonesia | AY651377 |
9 | A/goose/Fujian/bb/2003 | Goose | 2003 | China | DQ997402 |
10 | A/Beijing/01/2003 | People | 2003 | China | EF587280 |
11 | A/duck/Hunan/1386/2003 | Duck | 2003 | China | CY029310 |
12 | A/Thailand/Kan353/2004 2004 | People | 2004 | Thailand | EF541446 |
13 | A/duck/Guangxi/351/2004 | Duck | 2004 | China | DQ320943 |
14 | A/Vietnam/CL100/2004 | People | 2004 | Vietnam | DQ492986 |
15 | A/duck/Vietnam/48/2004 2004 | Duck | 2004 | Vietnam | DQ492944 |
16 | A/Cambodia/P0322095/2005 | People | 2005 | cambodia divide | HQ200461 |
17 | A/Indonesia/239H/2005 | People | 2005 | Indonesia | EU146665 |
18 | A/black-headed goose/Qinghai/1/2005 | Goose | 2005 | China | DQ100567 |
19 | A/Anhui/2/2005 | People | 2005 | China | CY060147 |
20 | A/duck/Guiyang/3834/2005 | Duck | 2005 | China | EF124151 |
21 | A/chicken/Lang Son/200/2005 | Chicken | 2005 | Vietnam | GU186717 |
22 | A/goose/Vietnam/3/05 2005 | Goose | 2005 | Vietnam | DQ366317 |
23 | A/duck/Vietnam/286/2005 | Duck | 2005 | Vietnam | DQ492958 |
24 | A/crested myna/Hong Kong/540/2006 | Bird | 2006 | Hong Kong | EF12405 |
25 | A/Indonesia/535H/2006 | People | 2006 | Indonesia | EU146756 |
26 | A/chicken/Shandong/A-10/2006 | Chicken | 2006 | China | HM172139 |
27 | A/chicken/shanxi/10/2006 | Chicken | 2006 | China | HM172135.1 |
28 | A/chicken/shanxi/2/2006 | Chicken | 2006 | China | DQ914817 |
Maybe you are interested!
-
 Phylogenetic Tree Showing Relationships of Vsv Species in 5 Study Samples A1, A2, A3, A4 and A5.
Phylogenetic Tree Showing Relationships of Vsv Species in 5 Study Samples A1, A2, A3, A4 and A5. -
 Car body electrical practice - 8
zt2i3t4l5ee
zt2a3gs
zt2a3ge
zc2o3n4t5e6n7ts
If the voltage is out of specification, replace the wire or connector.
If the voltage is within specification, install the front fog light relay and follow step 5.
Step 5 Check the front fog light switch
- Remove the D4 connector of the fog light switch
- Use a multimeter to measure the resistance of the front fog light switch.
Measurement location
Condition
Standard
D4-3 (BFG) -D4-4 (LFG)
Light switchFront Fog OFF
>10kΩ
D4-3 (BFG) -D4-4 (LFG)
Front fog light switchON
<1 Ω
- Standard resistor
D4 connector is located on the combination switch assembly.
If the resistance is out of specification, replace the combination switch (the fog light switch is located in the combination switch).
If the resistance is within specification, follow step 6.
Step 6 Check wiring and connectors (front fog light relay-light selector switch)
- Disconnect connector D4 of the combination switch assembly
- Use a voltmeter to measure the voltage value of jack D4 on the wire side.
Measurement location
Control modecontrol
Standard
D4-3 (BFG) - (-) AQ
TAIL
11 to 14 V
D4 connector for the wiring of the combination switch assembly
If the voltage does not meet the standard, replace the wire or connector.
If the voltage is within standard, there may have been an error in the previous measurements.
Step 7 Check the front fog lights
- Remove the front fog light electrical connector.
- Supply battery voltage to the fog lamp terminals
Jack 8, B9 of front fog lamp on the electrical side
blind first.
Power supply location
Terms and Conditions
Battery positive terminal - Terminal 2Battery negative terminal - Terminal 1
Fog lightsbefore morning
- If the light does not come on, replace the bulb.
If the light is on, re-plug the jack and continue to step 8.
Step 8 Check wiring and connectors (relay and front fog lights)
- Disconnect the B8 and B9 connectors of the front fog lights.
- Use a voltmeter to measure voltage at the following locations:
Measurement location
Switch location
Terms and Conditions
B8-2 - (-) AQ
Electric lock ON TAIL size switchFog switch ON
11 to 14 V
B9-2 - (-) AQ
Electric lock ONTAIL size switch Fog switch ON
11 to 14 V
B8 and B9 connectors on the front fog lamp wiring side
Voltage is not up to standard, repair or replace the jack. If up to standard, there may have been an error in the measurement process.
2.2.4. Procedure for removing, installing and adjusting fog lights 1. Procedure for removing
- Remove the front inner ear pads
Use a screwdriver to remove the 3 screws and remove the front part of the front inner ear liner
-Remove the fog light assembly
+ Disconnect the connector.
+ Use a screwdriver to remove 3 screws to remove the fog light cover
2. Installation sequence
-Rotate the fog lamp bulb in the direction indicated by the arrow as shown in the figure and remove the fog lamp from the fog lamp assembly.
-Rotate the fog light bulb in the direction indicated by the arrow as shown in the figure and install the light into the fog light assembly.
- Use a screwdriver to install the fog light cover
-Install the electrical connector
Attention: Be careful not to damage the plastic thread on the lamp assembly.
- Install the front inner ear pads
Use a screwdriver to install the front inner bumper with 3 screws.
3. Prepare the vehicle to adjust the fog light convergence. Prepare the vehicle:
- Make sure there is no damage or deformation to the vehicle body around the fog lights.
- Add fuel to the fuel tank
- Add oil to standard level.
- Add engine coolant to standard level.
- Inflate the tire to standard pressure.
- Place spare tire, tools and jack in original design position
- Do not leave any load in the luggage compartment.
- Let a person weighing about 75 kg sit in the driver's seat.
4. Prepare to check the fog light convergence
a/ Prepare the vehicle status as follows:
- Place the car in a dark enough place to see the lines. The lines are the dividing line, below which the light from the fog lights can be seen but above which it cannot.
- Place the car perpendicular to the wall.
- Keep a distance of 7.62 m between the center of the fog lamp and the wall.
- Park the car on level ground.
- Press the car down a few times to stabilize the suspension.
Note: A distance of approximately 7.62 m is required between the vehicle (fog lamp center) and the wall to adjust the convergence correctly. If the distance of 7.62 m cannot be achieved, set the correct distance of 3 m to check and adjust the fog lamp convergence. (Since the target area varies with the distance, please follow the instructions as shown in the figure.)
b/ Prepare a piece of thick white paper about 2 m high and 4 m wide to use as a screen.
c/ Draw a vertical line through the center of the screen (line V).
d/ Set the screen as shown in the picture. Note:
- Keep the screen perpendicular to the ground.
- Align the V line on the screen with the center of the vehicle.
e/Draw the reference lines (H, V LH and V RH lines) on the screen as shown in the figure.HINT:
Mark the center of the fog lamp on the screen. If the center mark cannot be seen on the fog lamp, use the center of the fog lamp or the manufacturer's name mark on the fog lamp as the center mark.
H line (fog light height):
Draw a line across the screen so that it passes through the center mark. Line H should be at the same height as the center mark of the fog light bulb.
Line V LH, V RH (center mark position of left fog lamp LH and right fog lamp RH):
Draw two lines so that they intersect line H at the center marks.
5. Check the fog light convergence
a/ Cover the fog lamp or remove the connector of the other side fog lamp to prevent light from the unchecked fog lamp from affecting the fog lamp convergence test.
b/ Start the engine.
c/ Turn on the fog lights and make sure that the dividing line is outside the standard area as shown in the drawing.
6. Adjust the fog light convergence
Use a screwdriver to adjust the fog light to the standard area by turning the toe adjustment screw.
Note: If the screw is adjusted too far, loosen it and then tighten it again, so that the last rotation of the light adjustment screw is clockwise.
3. Self-study questions
1. Describe the operating principle of the lighting system with automatic headlight function
2. Describe the operating principle of the lighting system with the function of rotating headlights when turning
3. Draw diagram and connect lighting system on Hyundai Porter car
4. Draw diagram and connect lighting system on Honda Accord 1992
5. Draw the lighting circuit on a 1993 Toyota Lexus
LESSON 3 MAINTENANCE AND REPAIR OF SIGNAL SYSTEM
I. IMPLEMENTATION GOAL
After completing this lesson, students will be able to:
- Distinguish between types of signals on cars
- Correctly describe common symptoms and suspected areas causing damage.
- Connecting signal circuits ensures technical requirements
- Disassemble, install, check, maintain and repair the signal system to ensure technical requirements.
- Ensure safety in work and industrial hygiene
II. LESSON CONTENT
1. General description
The signal system equipped on cars aims to create signals to notify other vehicles participating in traffic about the vehicle's operating status such as: stopping, parking, braking, reversing, turning...
Signals are used either by light such as headlamps, brake lights, turn signals….. or by sound such as horns, reverse music….
Just like the lighting system. A signal system circuit usually consists of: battery, fuse, wire, relay, electrical load and control switch. Only some switches of the signal system are on the combination switch. The switches of other signals are usually located in different locations such as in the gearbox or brake pedal……
2. Maintenance and repair
2.1. Turn signals and hazard lights
The installation location of the turn signal is shown in Figure 3.1. The turn signal control switch is located in the combination switch under the steering wheel. Turning this switch to the right or left will make the turn signal turn right or left.
The hazard light switch is used when the vehicle has a problem while participating in traffic. When the hazard light switch is turned on, all the turn signals on the vehicle will light up at a certain frequency. The hazard light switch is usually placed separately from the turn signal switch (some old cars integrate the hazard and turn signal switches on the same combination switch cluster).
Figure 3.1 Turn signal switch Figure 3.2 Hazard switch
The part that generates the flashing frequency for the lights is called a turn signal relay. The turn signal relay usually has 3 terminals: B (positive power supply); E (negative power supply); L (providing the turn signal switch to distribute to the
lamp)
2.1.1. Circuit diagram
To generate the frequency for the turn signal, a turn signal relay is used in the turn signal circuit. The current from the turn signal relay will be sent to the turn signal switch assembly to distribute the current to the turn signal lights for the driver's purpose.
Figure 3.3. Schematic diagram of a turn signal circuit without a hazard switch
1. Battery; 2. Electric lock; 3. Turn signal relay; 4. Turn signal switch; 5. Turn signal lamp; 6. Turn signal lamp; 7. Hazard switch
Figure 3.4 Schematic diagram of turn signal circuit with hazard switch
1. Battery; 2. Combination switch cluster; 3. Turn signal;
4. Turn signal light; 5. Turn signal relay
Today's cars no longer use three-pin turn signal relays (B, L, E) but use eight-pin turn signal relays (figure 3.5) (pin number 8 is used for hazard lights).
For this type, the current supplying the turn signal lights is supplied directly from the turn signal relay to the lights.
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s3 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; vertical-align: 1pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -9pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s9 { color: #008000; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; te
Car body electrical practice - 8
zt2i3t4l5ee
zt2a3gs
zt2a3ge
zc2o3n4t5e6n7ts
If the voltage is out of specification, replace the wire or connector.
If the voltage is within specification, install the front fog light relay and follow step 5.
Step 5 Check the front fog light switch
- Remove the D4 connector of the fog light switch
- Use a multimeter to measure the resistance of the front fog light switch.
Measurement location
Condition
Standard
D4-3 (BFG) -D4-4 (LFG)
Light switchFront Fog OFF
>10kΩ
D4-3 (BFG) -D4-4 (LFG)
Front fog light switchON
<1 Ω
- Standard resistor
D4 connector is located on the combination switch assembly.
If the resistance is out of specification, replace the combination switch (the fog light switch is located in the combination switch).
If the resistance is within specification, follow step 6.
Step 6 Check wiring and connectors (front fog light relay-light selector switch)
- Disconnect connector D4 of the combination switch assembly
- Use a voltmeter to measure the voltage value of jack D4 on the wire side.
Measurement location
Control modecontrol
Standard
D4-3 (BFG) - (-) AQ
TAIL
11 to 14 V
D4 connector for the wiring of the combination switch assembly
If the voltage does not meet the standard, replace the wire or connector.
If the voltage is within standard, there may have been an error in the previous measurements.
Step 7 Check the front fog lights
- Remove the front fog light electrical connector.
- Supply battery voltage to the fog lamp terminals
Jack 8, B9 of front fog lamp on the electrical side
blind first.
Power supply location
Terms and Conditions
Battery positive terminal - Terminal 2Battery negative terminal - Terminal 1
Fog lightsbefore morning
- If the light does not come on, replace the bulb.
If the light is on, re-plug the jack and continue to step 8.
Step 8 Check wiring and connectors (relay and front fog lights)
- Disconnect the B8 and B9 connectors of the front fog lights.
- Use a voltmeter to measure voltage at the following locations:
Measurement location
Switch location
Terms and Conditions
B8-2 - (-) AQ
Electric lock ON TAIL size switchFog switch ON
11 to 14 V
B9-2 - (-) AQ
Electric lock ONTAIL size switch Fog switch ON
11 to 14 V
B8 and B9 connectors on the front fog lamp wiring side
Voltage is not up to standard, repair or replace the jack. If up to standard, there may have been an error in the measurement process.
2.2.4. Procedure for removing, installing and adjusting fog lights 1. Procedure for removing
- Remove the front inner ear pads
Use a screwdriver to remove the 3 screws and remove the front part of the front inner ear liner
-Remove the fog light assembly
+ Disconnect the connector.
+ Use a screwdriver to remove 3 screws to remove the fog light cover
2. Installation sequence
-Rotate the fog lamp bulb in the direction indicated by the arrow as shown in the figure and remove the fog lamp from the fog lamp assembly.
-Rotate the fog light bulb in the direction indicated by the arrow as shown in the figure and install the light into the fog light assembly.
- Use a screwdriver to install the fog light cover
-Install the electrical connector
Attention: Be careful not to damage the plastic thread on the lamp assembly.
- Install the front inner ear pads
Use a screwdriver to install the front inner bumper with 3 screws.
3. Prepare the vehicle to adjust the fog light convergence. Prepare the vehicle:
- Make sure there is no damage or deformation to the vehicle body around the fog lights.
- Add fuel to the fuel tank
- Add oil to standard level.
- Add engine coolant to standard level.
- Inflate the tire to standard pressure.
- Place spare tire, tools and jack in original design position
- Do not leave any load in the luggage compartment.
- Let a person weighing about 75 kg sit in the driver's seat.
4. Prepare to check the fog light convergence
a/ Prepare the vehicle status as follows:
- Place the car in a dark enough place to see the lines. The lines are the dividing line, below which the light from the fog lights can be seen but above which it cannot.
- Place the car perpendicular to the wall.
- Keep a distance of 7.62 m between the center of the fog lamp and the wall.
- Park the car on level ground.
- Press the car down a few times to stabilize the suspension.
Note: A distance of approximately 7.62 m is required between the vehicle (fog lamp center) and the wall to adjust the convergence correctly. If the distance of 7.62 m cannot be achieved, set the correct distance of 3 m to check and adjust the fog lamp convergence. (Since the target area varies with the distance, please follow the instructions as shown in the figure.)
b/ Prepare a piece of thick white paper about 2 m high and 4 m wide to use as a screen.
c/ Draw a vertical line through the center of the screen (line V).
d/ Set the screen as shown in the picture. Note:
- Keep the screen perpendicular to the ground.
- Align the V line on the screen with the center of the vehicle.
e/Draw the reference lines (H, V LH and V RH lines) on the screen as shown in the figure.HINT:
Mark the center of the fog lamp on the screen. If the center mark cannot be seen on the fog lamp, use the center of the fog lamp or the manufacturer's name mark on the fog lamp as the center mark.
H line (fog light height):
Draw a line across the screen so that it passes through the center mark. Line H should be at the same height as the center mark of the fog light bulb.
Line V LH, V RH (center mark position of left fog lamp LH and right fog lamp RH):
Draw two lines so that they intersect line H at the center marks.
5. Check the fog light convergence
a/ Cover the fog lamp or remove the connector of the other side fog lamp to prevent light from the unchecked fog lamp from affecting the fog lamp convergence test.
b/ Start the engine.
c/ Turn on the fog lights and make sure that the dividing line is outside the standard area as shown in the drawing.
6. Adjust the fog light convergence
Use a screwdriver to adjust the fog light to the standard area by turning the toe adjustment screw.
Note: If the screw is adjusted too far, loosen it and then tighten it again, so that the last rotation of the light adjustment screw is clockwise.
3. Self-study questions
1. Describe the operating principle of the lighting system with automatic headlight function
2. Describe the operating principle of the lighting system with the function of rotating headlights when turning
3. Draw diagram and connect lighting system on Hyundai Porter car
4. Draw diagram and connect lighting system on Honda Accord 1992
5. Draw the lighting circuit on a 1993 Toyota Lexus
LESSON 3 MAINTENANCE AND REPAIR OF SIGNAL SYSTEM
I. IMPLEMENTATION GOAL
After completing this lesson, students will be able to:
- Distinguish between types of signals on cars
- Correctly describe common symptoms and suspected areas causing damage.
- Connecting signal circuits ensures technical requirements
- Disassemble, install, check, maintain and repair the signal system to ensure technical requirements.
- Ensure safety in work and industrial hygiene
II. LESSON CONTENT
1. General description
The signal system equipped on cars aims to create signals to notify other vehicles participating in traffic about the vehicle's operating status such as: stopping, parking, braking, reversing, turning...
Signals are used either by light such as headlamps, brake lights, turn signals….. or by sound such as horns, reverse music….
Just like the lighting system. A signal system circuit usually consists of: battery, fuse, wire, relay, electrical load and control switch. Only some switches of the signal system are on the combination switch. The switches of other signals are usually located in different locations such as in the gearbox or brake pedal……
2. Maintenance and repair
2.1. Turn signals and hazard lights
The installation location of the turn signal is shown in Figure 3.1. The turn signal control switch is located in the combination switch under the steering wheel. Turning this switch to the right or left will make the turn signal turn right or left.
The hazard light switch is used when the vehicle has a problem while participating in traffic. When the hazard light switch is turned on, all the turn signals on the vehicle will light up at a certain frequency. The hazard light switch is usually placed separately from the turn signal switch (some old cars integrate the hazard and turn signal switches on the same combination switch cluster).
Figure 3.1 Turn signal switch Figure 3.2 Hazard switch
The part that generates the flashing frequency for the lights is called a turn signal relay. The turn signal relay usually has 3 terminals: B (positive power supply); E (negative power supply); L (providing the turn signal switch to distribute to the
lamp)
2.1.1. Circuit diagram
To generate the frequency for the turn signal, a turn signal relay is used in the turn signal circuit. The current from the turn signal relay will be sent to the turn signal switch assembly to distribute the current to the turn signal lights for the driver's purpose.
Figure 3.3. Schematic diagram of a turn signal circuit without a hazard switch
1. Battery; 2. Electric lock; 3. Turn signal relay; 4. Turn signal switch; 5. Turn signal lamp; 6. Turn signal lamp; 7. Hazard switch
Figure 3.4 Schematic diagram of turn signal circuit with hazard switch
1. Battery; 2. Combination switch cluster; 3. Turn signal;
4. Turn signal light; 5. Turn signal relay
Today's cars no longer use three-pin turn signal relays (B, L, E) but use eight-pin turn signal relays (figure 3.5) (pin number 8 is used for hazard lights).
For this type, the current supplying the turn signal lights is supplied directly from the turn signal relay to the lights.
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s3 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; vertical-align: 1pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -9pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s9 { color: #008000; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; te -
 Illustration of Teaching Organization Model Using M-Learning
Illustration of Teaching Organization Model Using M-Learning -
 Sequence Diagram of Accounting for Other Operating Income
Sequence Diagram of Accounting for Other Operating Income -
 Study on natural regeneration characteristics of Lim xet tree species Peltophorum tonkinensis A.Chev in Lam Binh district, Tuyen Quang province - 13
Study on natural regeneration characteristics of Lim xet tree species Peltophorum tonkinensis A.Chev in Lam Binh district, Tuyen Quang province - 13

STT
International trade | Species battery | Capacity set up | Place of isolation | S bank registration gene | |
29 | A/Anhui/T2/2006 2006 | People | 2006 | China | EU008577 |
30 | A/Hunan/1/2006 | People | 2006 | China | CY060173 |
31 | A/Shanghai/1/2006 | People | 2006 | China | AB462298 |
32 | A/duck/Guangxi/150/2006 | Duck | 2006 | China | EF124078 |
33 | A/chicken/East Java/UT6045/2007 | Chicken | 2007 | Indonesia | GQ122542 |
34 | A/Indonesia/CDC1046/2007 | People | 2007 | Indonesia | CY019411 |
35 | A/duck/Laos/P0117/2007 | Duck | 2007 | Laos | CY040921 |
36 | A/Muscovy duck/Ca Mau/07- April 2007 | Goose | 2007 | Vietnam | GU186746 |
37 | A/duck/India/Tr-NIV4396/2008 | Duck | 2008 | India | CY046105 |
38 | A/chicken/Bhutan/248006/2010 | Chicken | 2010 | Bhutan | CY066007 |
39 | A/mallard/Korea/1195/2010 | Duck God | 2010 | Korea | HQ695916 |
40 | A/Hong Kong/6841/2010 | People | 2010 | Hong Kong | HQ636463 |
41 | A/chicken/Thailand/ICRC- 7356/2010 | Chicken | 2010 | Thailand | HM590792 |
Based on the analysis of the gene family tree, we can conclude that the avian influenza A/H5N1 virus strains of clade 7 in Vietnam have high similarity and are arranged in a group with each other, and in a group with the influenza A/H5N1 virus strains of China. This analysis results in the M gene of the influenza A/H5N1 virus of clade 7 in Vietnam and China having the same origin. To see more clearly the genetic differences at the nucleotide and amino acid levels, we compared the genes of the influenza A/H5N1 virus strains of Vietnam with some influenza A/H5N1 viruses of China. The results are shown in Fig.
Table 3.8.
The level of genetic difference in M gene between Vietnamese A/H5N1 clade 7 influenza virus strains is almost negligible, with the maximum difference in nucleotide level being 1% and amino acid level being 0.6%.
The level of difference between the Vietnamese A/H5N1 influenza virus strains belonging to clade 7 and the strains belonging to clade 7 in China is also very low. The minimum difference
The genetic diversity at the nucleotide and amino acid levels between the viruses of these two groups was 3.0±0.1% and 3.3±0.2%, respectively (virus A chicken/Shanxi/2/2006).
Comparing the genetic differences between the influenza virus strains of the clade we isolated with those of other clade viruses on the basis of genes, we found that the differences in nucleotides and amino acids were not much, only about 3.6% in nucleotides and 2-4% in amino acids. Only the two strains A goose Vietnam 3 0 and A HongKong 1 6 9 belonging to clade 0 had quite large differences: the difference in nucleotide levels was up to 10% and the difference in amino acids was up to 7%. Another AH N1 virus strain also belonging to clade 0, the A Goose Guangdong 1 96 strain, considered the ancestor of highly pathogenic AH N1 influenza virus strains, had genetic differences with the virus strains belonging to clade 0.
In Vietnam, the nucleotide level was .3±0.2% (.2-.6%) and the amino acid level was 4.2±0.2% (4.1-4.4%).
In summary, the results of phylogenetic analysis and genetic differences analysis based on the H, N1 genes of the A/H5N1 clade 7 influenza virus strains in Vietnam show that the virus strains belonging to clade 7 in Vietnam have high similarity with each other, proving that they have the same origin, and are similar to the AH N1 virus strains also belonging to the clade isolated in China.
From the above results, it can be concluded that the AH N1 influenza virus strains belonging to clade 7 in Vietnam have the same origin as the viruses belonging to clade 7 in China, and have entered Vietnam, not viruses that have evolved from highly pathogenic AH N1 influenza virus strains that previously caused disease in poultry (2003-2008) in Vietnam. The phenomenon of virus strains belonging to the clade discovered in the border area in 2008, along with the appearance of many other clades in Vietnam such as clade 2.3.4 and 2.3.2 in Vietnam, which are very similar to the AH N1 influenza virus strains circulating and causing disease in Chinese poultry, shows that the risk of infiltration and emergence of A/H5N1 influenza virus strains from outside into Vietnam is very high, so it is necessary to strengthen surveillance to detect early AH N1 influenza viruses that may invade or are circulating in Vietnamese poultry.
Table 3.8. Comparison of genetic differences in the M gene of 5 A/H5N1 virus strains belonging to clade 7
remember some reference strains
Species symbol
Strain name i 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15
Amino acids (%)
1
A/Chicken/Vietnam/NCVD-016/2008 | 0.6 | 0.3 | 0.3 | 0.3 | 1.6 | 1.3 | 3.5 | 1.9 | 7.4 | 1.9 | 2.1 | 3.8 | 4.4 | 7.4 | ||
2 | A/chicken/Vietnam/NCVD-093/2008 | 1.0 | 0.3 | 0.3 | 0.3 | 1.6 | 1.3 | 3.5 | 1.9 | 7.4 | 1.9 | 1.8 | 3.8 | 4.4 | 7.4 | |
3 | A/chicken/Vietnam/NCVD-03/2008 | 0.9 | 0.3 | 0.0 | 0.0 | 1.3 | 0.9 | 3.2 | 1.6 | 7.1 | 1.6 | 1.8 | 3.5 | 4.1 | 7.1 | |
4 | A/chicken/Vietnam/NCVD-04/2008 | 0.9 | 0.3 | 0.2 | 0.0 | 1.3 | 0.9 | 3.2 | 1.6 | 7.1 | 1.6 | 1.8 | 3.5 | 4.1 | 7.1 | |
5 | A/chicken/Vietnam/NCVD-05/2008 | 0.9 | 0.3 | 0.2 | 0.0 | 1.3 | 0.9 | 3.2 | 1.6 | 7.1 | 1.6 | 1.8 | 3.5 | 4.1 | 7.1 | |
6 | A/chicken/Shandong/A-10/2006 | 1.3 | 1.3 | 1.2 | 1.2 | 1.2 | 0.9 | 3.8 | 1.6 | 7.4 | 1.6 | 1.8 | 3.5 | 4.4 | 7.4 | |
7 | A/chicken/Shanxi/10/2006 | 0.8 | 0.9 | 0.8 | 0.8 | 0.8 | 0.8 | 4.1 | 1.6 | 7.4 | 1.6 | 1.8 | 3.5 | 4.4 | 7.4 | |
8 | A/chicken/Shanxi/2/2006 | 3.1 | 3.1 | 2.8 | 2.9 | 2.9 | 2.8 | 3.0 | 3.2 | 7.4 | 3.2 | 3.5 | 5.1 | 5.8 | 7.4 | |
9 | A/Beijing/01/2003_2003 | 2.4 | 2.4 | 2.3 | 2.3 | 2.3 | 1.7 | 2.1 | 3.5 | 5.8 | 0.0 | 0.7 | 1.9 | 2.8 | 5.8 | |
10 | A/goose/Vietnam/3/05 | 10.0 | 10.1 | 9.7 | 9.7 | 9.7 | 10.0 | 10.4 | 9.3 | 8.7 | 5.8 | 5.7 | 5.8 | 3.8 | 0.0 | |
11 | A/Anhui/2/2005 | 3.0 | 3.1 | 3.0 | 3.0 | 3.0 | 2.4 | 2.7 | 4.0 | 0.9 | 8.1 | 0.7 | 1.9 | 2.8 | 5.8 | |
12 | A/blackheadedgoose/Qinghai/January 2005 | 3.2 | 3.2 | 3.2 | 3.2 | 3.2 | 2.6 | 2.7 | 4.0 | 1.4 | 9.3 | 1.3 | 2.1 | 3.2 | 5.7 | |
13 | A/goose/Fujian/bb/2003 | 4.0 | 4.1 | 3.8 | 3.8 | 3.8 | 3.4 | 3.7 | 4.4 | 2.1 | 8.1 | 2.0 | 2.3 | 2.8 | 5.8 | |
14 | A/Goose/Guangdong/1/96 | 5.5 | 5.6 | 5.2 | 5.2 | 5.2 | 5.0 | 5.3 | 5.7 | 3.5 | 6.0 | 3.4 | 3.8 | 3.2 | 3.8 | |
15 | A/Hong_Kong/156/97 | 10.7 | 10.8 | 10.5 | 10.5 | 10.5 | 10.7 | 11.1 | 9.3 | 9.4 | 1.3 | 8.8 | 9.8 | 8.8 | 7.5 |
Nucleotides (%)
83
3.3. Determination of biological characteristics of A/H5N1 clade 7 virus isolated in Vietnam (A/chicken/Vietnam/NCVD-016/2008)
3.3.1. Chicken breed adaptation (EID 50 specification )
Influenza viruses in general and highly pathogenic avian influenza A/H N1 viruses have the ability to grow well on chicken embryos, so 9-10 day old chicken embryos are often used to isolate highly pathogenic avian influenza viruses in diagnostic work as well as to maintain virus strains. However, avian influenza virus strains have different characteristics when multiplying on egg embryos, as shown by embryo infectivity, embryo lethality and optimal titer of virus when multiplying on egg embryos. The differences in embryo lethality time and optimal titer of different virus strains are different. Low pathogenic viruses usually kill egg embryos more slowly and can obtain higher virus concentrations than highly pathogenic influenza viruses and vice versa.
To determine the adaptability of H5N1 clade avian influenza virus strains
isolated, we chose strain A/Chicken/Vietnam/NCVD-016/2008 as the representative strain because this strain can be considered close to the original strain of the clade from China.
Embryos were obtained from healthy chicken flocks that had not been vaccinated against H N1 avian influenza, and were not naturally infected with avian influenza virus to avoid the impact (if any) of antibodies from vaccinated hens (in the yolk of the embryos). Eggs were purchased at 8 days of age and incubated for 1 day to stabilize in the laboratory before being used for the experiment to determine the adaptation of the influenza virus strain A/Chicken/Vietnam/NCVD-016 2008 on chicken embryos.
The influenza virus strain A/Chicken/Vietnam/NCVD-016/2008 was divided into many small tubes after isolation and kept at -86 0 C. Before conducting the experiment, the virus suspension was checked for sterility according to the routine and was mixed into many concentrations from 10 -1 to 10 -8 with PBS buffer solution at pH 7.2. Egg embryos were infected with 6 dilutions from 10 -3 to 10 -8 into the allantoic cavity of the egg embryo, each concentration infected
embryos with a dose of 100µl embryos, continue incubation at 37 0 C, monitor twice a day
about 8 hours within 4 days, embryos that died before 24 hours. In this experiment, 3 embryos injected with PBS solution were used as negative controls and incubated under the same conditions.
The embryos that died during the monitoring period were cooled at 2-8 0 C for 24 hours to restore blood vessels and the eggs were dissected to harvest the tissue fluid. The embryos that survived after the monitoring period (including negative controls) were all collected for tissue fluid and tested by HA reaction to determine the virus infection of the embryos and determine the EID 50 index according to the Reed-Muench formula.
Before infection, we tested the sterility of the virus tubes on the following media: meat broth, BHI, regular agar, blood agar, Sabauraud and anaerobic liver broth, with the aim of determining for sure that the death of the embryos was caused by the influenza virus. We checked the embryos twice a day, 8 hours apart, to record the details of the time of death of the embryos. The results of the time of death of the embryos are presented in Table 3.9.
The results of determining the infectivity of chicken embryos and the EID 50 index of the virus strain A/Chicken/Vietnam/NCVD-016/2008 are presented in Table 3.10.
Table 3.9. Time to lethality monitoring
TN batch
Number of infected embryos | Tracking dead embryos | Total number of dead embryos | Rate (%) | ||||
0-24 h | 25-48 h | 49-72 hours | 73-96h | ||||
1 | 30 | 0 | 10 | 5 | 5 | 20 | 67 |
2 | 30 | 0 | 10 | 5 | 4 | 19 | 63 |
3 | 30 | 0 | 10 | 5 | 4 | 19 | 63 |
The results showed that: no embryos died before 24 hours, the virus started killing embryos within 2 days after infection. This is also consistent with our daily observations when isolating samples containing avian influenza H N1 virus on egg embryos, and also consistent with previous studies (Easterday
et al., 199; Wanasawaeng et al., 2009) observed that highly pathogenic influenza viruses usually kill embryonated eggs within 48 hours post-infection (hpi) of the urothelial sinus.
Table 3.10. Results of monitoring the survival/death ratio of eggs when infected with virus A/Chicken/Vietnam/NCVD-016/2008
Experimental times
Dilution titer | Number of infected embryos | Number of infected embryos | Number of uninfected embryos | Accumulated number | Virus infection rate (%) A (A+B) × 100 | |||
Viral infection (A) | Not infected (B) | Total (A+B) | ||||||
First time | 10 -3 | 5 | 5 | 0 | 20 | 0 | 20 | 100 |
10 -4 | 5 | 5 | 0 | 15 | 0 | 15 | 100 | |
10 -5 | 5 | 5 | 0 | 10 | 0 | 10 | 100 | |
10 -6 | 5 | 4 | 1 | 5 | 1 | 6 | 83.3 | |
10 -7 | 5 | 1 | 4 | 1 | 5 | 6 | 16.7 | |
10 -8 | 5 | 0 | 5 | 0 | 10 | 10 | 0 | |
DC (-) | 3 | 0 | 3 | |||||
EID 50 /0.1ml | 10 7 (according to the eed-Muench formula) | |||||||
2nd time | 10 -3 | 5 | 5 | 0 | 19 | 0 | 19 | 100 |
10 -4 | 5 | 5 | 0 | 14 | 0 | 14 | 100 | |
10 -5 | 5 | 5 | 0 | 9 | 0 | 9 | 100 | |
10 -6 | 5 | 3 | 2 | 4 | 1 | 5 | 80.0 | |
10 -7 | 5 | 1 | 4 | 1 | 5 | 6 | 16.7 | |
10 -8 | 5 | 0 | 5 | 0 | 10 | 10 | 0 | |
DC (-) | 3 | 0 | 3 | |||||
EID 50 /0.1 ml | 10 6.9 (according to the eed-Muench formula) | |||||||
3rd time | 10 -3 | 5 | 5 | 0 | 19 | 0 | 19 | 100 |
10 -4 | 5 | 5 | 0 | 14 | 0 | 14 | 100 | |
10 -5 | 5 | 5 | 0 | 9 | 0 | 9 | 100 | |
10 -6 | 5 | 3 | 2 | 4 | 1 | 5 | 80.0 | |
10 -7 | 5 | 1 | 4 | 1 | 5 | 6 | 16.7 | |
10 -8 | 5 | 0 | 5 | 0 | 10 | 10 | 0 | |
DC (-) | 3 | 0 | 3 | 0 | ||||
EID 50 /0.1 ml | 10 6.9 (according to the eed-Muench formula) | |||||||
- At a dilution concentration of 10-3 , all embryos died within 40 hours after infection.