consists of a number of alkaline amino acids encoded by an oligonucleotide sequence, which is the protease cleavage site, which is the region that determines the virulence of the virus (Bosch et al., 1981; Gambotto et al., 2008). The HA protein has a molecular mass of approximately 63,103 Da (if not glycosylated) and .103 Da (if glycosylated, in which HA1 is 48,103 Da and HA2 is 29,103 Da) (Keawcharoen et al., 2000; Luong and Palese, 1992).
- Segment 5 (NP gene) is about 1 6 bp in size, encoding the synthesis of nucleoprotein (NP) - a component of the transcription complex, responsible for transporting NA between the nucleus and the host cell cytoplasm. NP is a glycosylated protein, with antigenic properties expressed by viral groups, existing in virus particles in association with each NA segment, with a calculated molecular mass of about 6.103 Da (in fact 0 - 60.103 Da) (Luong and Palese, 1992; urphy and Webster, 1996; ӧmer-Oberdӧrfer et al., 2008).
- Segment 6 (NA gene) is an antigenic gene of the virus, with a length that varies with each strain of influenza A virus (A H6N2 is 1413 bp, AH N1 varies from about 130 - 1410 bp) (Le Thanh Hoa, 2004). This is the gene that encodes the synthesis of the NA protein, the surface antigen of the capsid of the virus, with a calculated molecular mass of about 0.103 Da (in fact, 0 - 60.103 Da). Molecular studies of the NA gene of the influenza virus show that the '- end of this gene (or the N-terminal part of the NA polypeptide) is highly variable and complex among influenza A virus strains, this change is related to the adaptation and pathogenicity of the influenza virus on many different hosts (Castrucci and Kawaoka, 1993; Baigent et al. Cauley, 2001; Uiprasertkul et al. 2000). The characteristic mutation of the NA gene in influenza A virus is the phenomenon of sliding-deletion of a gene segment of nucleotide, then 60 nucleotides, making the original length of NA(N1) of 1410 bp reduced to 130 bp (Castrucci and Kawaoka, 1993; Le Thanh Hoa, 2006; Kaewcharoen et al., 200).
The gene segments, NP and NS, encode the synthesis of different functional viral proteins, which are relatively stable in length among influenza A virus strains, including:
- Segment 7 (gene) is about 102 bp in size, encoding the matrix protein (matrix protein - ) of the virus (consisting of two subunits 1 and 2 created by different reading frames of the same NA segment), together with HA and NA, there are about 3000 P molecules on the surface of the influenza A virus capsid, which has a surface interaction relationship with hemagglutinin (Scholtissek et al., 2002). Protein 1 is a matrix protein, the main component of the virus that functions to encapsulate NA to form the NP complex and participate in the "budding" process of the virus (Luong and Palese, 1993; Urphy and Webster, 1996; Basler, 2000). Protein 2 is a small polypeptide chain, with a calculated molecular weight of 11.103 Da (actually 1.103 Da), a membrane-permeable protein - ion channel necessary for viral infectivity, responsible for "coating" the virus to present the host cell cytoplasmic genome during host infection (Scholtissek et al., 2002).
Maybe you are interested!
-
 Car body electrical practice - 8
zt2i3t4l5ee
zt2a3gs
zt2a3ge
zc2o3n4t5e6n7ts
If the voltage is out of specification, replace the wire or connector.
If the voltage is within specification, install the front fog light relay and follow step 5.
Step 5 Check the front fog light switch
- Remove the D4 connector of the fog light switch
- Use a multimeter to measure the resistance of the front fog light switch.
Measurement location
Condition
Standard
D4-3 (BFG) -D4-4 (LFG)
Light switchFront Fog OFF
>10kΩ
D4-3 (BFG) -D4-4 (LFG)
Front fog light switchON
<1 Ω
- Standard resistor
D4 connector is located on the combination switch assembly.
If the resistance is out of specification, replace the combination switch (the fog light switch is located in the combination switch).
If the resistance is within specification, follow step 6.
Step 6 Check wiring and connectors (front fog light relay-light selector switch)
- Disconnect connector D4 of the combination switch assembly
- Use a voltmeter to measure the voltage value of jack D4 on the wire side.
Measurement location
Control modecontrol
Standard
D4-3 (BFG) - (-) AQ
TAIL
11 to 14 V
D4 connector for the wiring of the combination switch assembly
If the voltage does not meet the standard, replace the wire or connector.
If the voltage is within standard, there may have been an error in the previous measurements.
Step 7 Check the front fog lights
- Remove the front fog light electrical connector.
- Supply battery voltage to the fog lamp terminals
Jack 8, B9 of front fog lamp on the electrical side
blind first.
Power supply location
Terms and Conditions
Battery positive terminal - Terminal 2Battery negative terminal - Terminal 1
Fog lightsbefore morning
- If the light does not come on, replace the bulb.
If the light is on, re-plug the jack and continue to step 8.
Step 8 Check wiring and connectors (relay and front fog lights)
- Disconnect the B8 and B9 connectors of the front fog lights.
- Use a voltmeter to measure voltage at the following locations:
Measurement location
Switch location
Terms and Conditions
B8-2 - (-) AQ
Electric lock ON TAIL size switchFog switch ON
11 to 14 V
B9-2 - (-) AQ
Electric lock ONTAIL size switch Fog switch ON
11 to 14 V
B8 and B9 connectors on the front fog lamp wiring side
Voltage is not up to standard, repair or replace the jack. If up to standard, there may have been an error in the measurement process.
2.2.4. Procedure for removing, installing and adjusting fog lights 1. Procedure for removing
- Remove the front inner ear pads
Use a screwdriver to remove the 3 screws and remove the front part of the front inner ear liner
-Remove the fog light assembly
+ Disconnect the connector.
+ Use a screwdriver to remove 3 screws to remove the fog light cover
2. Installation sequence
-Rotate the fog lamp bulb in the direction indicated by the arrow as shown in the figure and remove the fog lamp from the fog lamp assembly.
-Rotate the fog light bulb in the direction indicated by the arrow as shown in the figure and install the light into the fog light assembly.
- Use a screwdriver to install the fog light cover
-Install the electrical connector
Attention: Be careful not to damage the plastic thread on the lamp assembly.
- Install the front inner ear pads
Use a screwdriver to install the front inner bumper with 3 screws.
3. Prepare the vehicle to adjust the fog light convergence. Prepare the vehicle:
- Make sure there is no damage or deformation to the vehicle body around the fog lights.
- Add fuel to the fuel tank
- Add oil to standard level.
- Add engine coolant to standard level.
- Inflate the tire to standard pressure.
- Place spare tire, tools and jack in original design position
- Do not leave any load in the luggage compartment.
- Let a person weighing about 75 kg sit in the driver's seat.
4. Prepare to check the fog light convergence
a/ Prepare the vehicle status as follows:
- Place the car in a dark enough place to see the lines. The lines are the dividing line, below which the light from the fog lights can be seen but above which it cannot.
- Place the car perpendicular to the wall.
- Keep a distance of 7.62 m between the center of the fog lamp and the wall.
- Park the car on level ground.
- Press the car down a few times to stabilize the suspension.
Note: A distance of approximately 7.62 m is required between the vehicle (fog lamp center) and the wall to adjust the convergence correctly. If the distance of 7.62 m cannot be achieved, set the correct distance of 3 m to check and adjust the fog lamp convergence. (Since the target area varies with the distance, please follow the instructions as shown in the figure.)
b/ Prepare a piece of thick white paper about 2 m high and 4 m wide to use as a screen.
c/ Draw a vertical line through the center of the screen (line V).
d/ Set the screen as shown in the picture. Note:
- Keep the screen perpendicular to the ground.
- Align the V line on the screen with the center of the vehicle.
e/Draw the reference lines (H, V LH and V RH lines) on the screen as shown in the figure.HINT:
Mark the center of the fog lamp on the screen. If the center mark cannot be seen on the fog lamp, use the center of the fog lamp or the manufacturer's name mark on the fog lamp as the center mark.
H line (fog light height):
Draw a line across the screen so that it passes through the center mark. Line H should be at the same height as the center mark of the fog light bulb.
Line V LH, V RH (center mark position of left fog lamp LH and right fog lamp RH):
Draw two lines so that they intersect line H at the center marks.
5. Check the fog light convergence
a/ Cover the fog lamp or remove the connector of the other side fog lamp to prevent light from the unchecked fog lamp from affecting the fog lamp convergence test.
b/ Start the engine.
c/ Turn on the fog lights and make sure that the dividing line is outside the standard area as shown in the drawing.
6. Adjust the fog light convergence
Use a screwdriver to adjust the fog light to the standard area by turning the toe adjustment screw.
Note: If the screw is adjusted too far, loosen it and then tighten it again, so that the last rotation of the light adjustment screw is clockwise.
3. Self-study questions
1. Describe the operating principle of the lighting system with automatic headlight function
2. Describe the operating principle of the lighting system with the function of rotating headlights when turning
3. Draw diagram and connect lighting system on Hyundai Porter car
4. Draw diagram and connect lighting system on Honda Accord 1992
5. Draw the lighting circuit on a 1993 Toyota Lexus
LESSON 3 MAINTENANCE AND REPAIR OF SIGNAL SYSTEM
I. IMPLEMENTATION GOAL
After completing this lesson, students will be able to:
- Distinguish between types of signals on cars
- Correctly describe common symptoms and suspected areas causing damage.
- Connecting signal circuits ensures technical requirements
- Disassemble, install, check, maintain and repair the signal system to ensure technical requirements.
- Ensure safety in work and industrial hygiene
II. LESSON CONTENT
1. General description
The signal system equipped on cars aims to create signals to notify other vehicles participating in traffic about the vehicle's operating status such as: stopping, parking, braking, reversing, turning...
Signals are used either by light such as headlamps, brake lights, turn signals….. or by sound such as horns, reverse music….
Just like the lighting system. A signal system circuit usually consists of: battery, fuse, wire, relay, electrical load and control switch. Only some switches of the signal system are on the combination switch. The switches of other signals are usually located in different locations such as in the gearbox or brake pedal……
2. Maintenance and repair
2.1. Turn signals and hazard lights
The installation location of the turn signal is shown in Figure 3.1. The turn signal control switch is located in the combination switch under the steering wheel. Turning this switch to the right or left will make the turn signal turn right or left.
The hazard light switch is used when the vehicle has a problem while participating in traffic. When the hazard light switch is turned on, all the turn signals on the vehicle will light up at a certain frequency. The hazard light switch is usually placed separately from the turn signal switch (some old cars integrate the hazard and turn signal switches on the same combination switch cluster).
Figure 3.1 Turn signal switch Figure 3.2 Hazard switch
The part that generates the flashing frequency for the lights is called a turn signal relay. The turn signal relay usually has 3 terminals: B (positive power supply); E (negative power supply); L (providing the turn signal switch to distribute to the
lamp)
2.1.1. Circuit diagram
To generate the frequency for the turn signal, a turn signal relay is used in the turn signal circuit. The current from the turn signal relay will be sent to the turn signal switch assembly to distribute the current to the turn signal lights for the driver's purpose.
Figure 3.3. Schematic diagram of a turn signal circuit without a hazard switch
1. Battery; 2. Electric lock; 3. Turn signal relay; 4. Turn signal switch; 5. Turn signal lamp; 6. Turn signal lamp; 7. Hazard switch
Figure 3.4 Schematic diagram of turn signal circuit with hazard switch
1. Battery; 2. Combination switch cluster; 3. Turn signal;
4. Turn signal light; 5. Turn signal relay
Today's cars no longer use three-pin turn signal relays (B, L, E) but use eight-pin turn signal relays (figure 3.5) (pin number 8 is used for hazard lights).
For this type, the current supplying the turn signal lights is supplied directly from the turn signal relay to the lights.
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s3 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; vertical-align: 1pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -9pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s9 { color: #008000; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; te
Car body electrical practice - 8
zt2i3t4l5ee
zt2a3gs
zt2a3ge
zc2o3n4t5e6n7ts
If the voltage is out of specification, replace the wire or connector.
If the voltage is within specification, install the front fog light relay and follow step 5.
Step 5 Check the front fog light switch
- Remove the D4 connector of the fog light switch
- Use a multimeter to measure the resistance of the front fog light switch.
Measurement location
Condition
Standard
D4-3 (BFG) -D4-4 (LFG)
Light switchFront Fog OFF
>10kΩ
D4-3 (BFG) -D4-4 (LFG)
Front fog light switchON
<1 Ω
- Standard resistor
D4 connector is located on the combination switch assembly.
If the resistance is out of specification, replace the combination switch (the fog light switch is located in the combination switch).
If the resistance is within specification, follow step 6.
Step 6 Check wiring and connectors (front fog light relay-light selector switch)
- Disconnect connector D4 of the combination switch assembly
- Use a voltmeter to measure the voltage value of jack D4 on the wire side.
Measurement location
Control modecontrol
Standard
D4-3 (BFG) - (-) AQ
TAIL
11 to 14 V
D4 connector for the wiring of the combination switch assembly
If the voltage does not meet the standard, replace the wire or connector.
If the voltage is within standard, there may have been an error in the previous measurements.
Step 7 Check the front fog lights
- Remove the front fog light electrical connector.
- Supply battery voltage to the fog lamp terminals
Jack 8, B9 of front fog lamp on the electrical side
blind first.
Power supply location
Terms and Conditions
Battery positive terminal - Terminal 2Battery negative terminal - Terminal 1
Fog lightsbefore morning
- If the light does not come on, replace the bulb.
If the light is on, re-plug the jack and continue to step 8.
Step 8 Check wiring and connectors (relay and front fog lights)
- Disconnect the B8 and B9 connectors of the front fog lights.
- Use a voltmeter to measure voltage at the following locations:
Measurement location
Switch location
Terms and Conditions
B8-2 - (-) AQ
Electric lock ON TAIL size switchFog switch ON
11 to 14 V
B9-2 - (-) AQ
Electric lock ONTAIL size switch Fog switch ON
11 to 14 V
B8 and B9 connectors on the front fog lamp wiring side
Voltage is not up to standard, repair or replace the jack. If up to standard, there may have been an error in the measurement process.
2.2.4. Procedure for removing, installing and adjusting fog lights 1. Procedure for removing
- Remove the front inner ear pads
Use a screwdriver to remove the 3 screws and remove the front part of the front inner ear liner
-Remove the fog light assembly
+ Disconnect the connector.
+ Use a screwdriver to remove 3 screws to remove the fog light cover
2. Installation sequence
-Rotate the fog lamp bulb in the direction indicated by the arrow as shown in the figure and remove the fog lamp from the fog lamp assembly.
-Rotate the fog light bulb in the direction indicated by the arrow as shown in the figure and install the light into the fog light assembly.
- Use a screwdriver to install the fog light cover
-Install the electrical connector
Attention: Be careful not to damage the plastic thread on the lamp assembly.
- Install the front inner ear pads
Use a screwdriver to install the front inner bumper with 3 screws.
3. Prepare the vehicle to adjust the fog light convergence. Prepare the vehicle:
- Make sure there is no damage or deformation to the vehicle body around the fog lights.
- Add fuel to the fuel tank
- Add oil to standard level.
- Add engine coolant to standard level.
- Inflate the tire to standard pressure.
- Place spare tire, tools and jack in original design position
- Do not leave any load in the luggage compartment.
- Let a person weighing about 75 kg sit in the driver's seat.
4. Prepare to check the fog light convergence
a/ Prepare the vehicle status as follows:
- Place the car in a dark enough place to see the lines. The lines are the dividing line, below which the light from the fog lights can be seen but above which it cannot.
- Place the car perpendicular to the wall.
- Keep a distance of 7.62 m between the center of the fog lamp and the wall.
- Park the car on level ground.
- Press the car down a few times to stabilize the suspension.
Note: A distance of approximately 7.62 m is required between the vehicle (fog lamp center) and the wall to adjust the convergence correctly. If the distance of 7.62 m cannot be achieved, set the correct distance of 3 m to check and adjust the fog lamp convergence. (Since the target area varies with the distance, please follow the instructions as shown in the figure.)
b/ Prepare a piece of thick white paper about 2 m high and 4 m wide to use as a screen.
c/ Draw a vertical line through the center of the screen (line V).
d/ Set the screen as shown in the picture. Note:
- Keep the screen perpendicular to the ground.
- Align the V line on the screen with the center of the vehicle.
e/Draw the reference lines (H, V LH and V RH lines) on the screen as shown in the figure.HINT:
Mark the center of the fog lamp on the screen. If the center mark cannot be seen on the fog lamp, use the center of the fog lamp or the manufacturer's name mark on the fog lamp as the center mark.
H line (fog light height):
Draw a line across the screen so that it passes through the center mark. Line H should be at the same height as the center mark of the fog light bulb.
Line V LH, V RH (center mark position of left fog lamp LH and right fog lamp RH):
Draw two lines so that they intersect line H at the center marks.
5. Check the fog light convergence
a/ Cover the fog lamp or remove the connector of the other side fog lamp to prevent light from the unchecked fog lamp from affecting the fog lamp convergence test.
b/ Start the engine.
c/ Turn on the fog lights and make sure that the dividing line is outside the standard area as shown in the drawing.
6. Adjust the fog light convergence
Use a screwdriver to adjust the fog light to the standard area by turning the toe adjustment screw.
Note: If the screw is adjusted too far, loosen it and then tighten it again, so that the last rotation of the light adjustment screw is clockwise.
3. Self-study questions
1. Describe the operating principle of the lighting system with automatic headlight function
2. Describe the operating principle of the lighting system with the function of rotating headlights when turning
3. Draw diagram and connect lighting system on Hyundai Porter car
4. Draw diagram and connect lighting system on Honda Accord 1992
5. Draw the lighting circuit on a 1993 Toyota Lexus
LESSON 3 MAINTENANCE AND REPAIR OF SIGNAL SYSTEM
I. IMPLEMENTATION GOAL
After completing this lesson, students will be able to:
- Distinguish between types of signals on cars
- Correctly describe common symptoms and suspected areas causing damage.
- Connecting signal circuits ensures technical requirements
- Disassemble, install, check, maintain and repair the signal system to ensure technical requirements.
- Ensure safety in work and industrial hygiene
II. LESSON CONTENT
1. General description
The signal system equipped on cars aims to create signals to notify other vehicles participating in traffic about the vehicle's operating status such as: stopping, parking, braking, reversing, turning...
Signals are used either by light such as headlamps, brake lights, turn signals….. or by sound such as horns, reverse music….
Just like the lighting system. A signal system circuit usually consists of: battery, fuse, wire, relay, electrical load and control switch. Only some switches of the signal system are on the combination switch. The switches of other signals are usually located in different locations such as in the gearbox or brake pedal……
2. Maintenance and repair
2.1. Turn signals and hazard lights
The installation location of the turn signal is shown in Figure 3.1. The turn signal control switch is located in the combination switch under the steering wheel. Turning this switch to the right or left will make the turn signal turn right or left.
The hazard light switch is used when the vehicle has a problem while participating in traffic. When the hazard light switch is turned on, all the turn signals on the vehicle will light up at a certain frequency. The hazard light switch is usually placed separately from the turn signal switch (some old cars integrate the hazard and turn signal switches on the same combination switch cluster).
Figure 3.1 Turn signal switch Figure 3.2 Hazard switch
The part that generates the flashing frequency for the lights is called a turn signal relay. The turn signal relay usually has 3 terminals: B (positive power supply); E (negative power supply); L (providing the turn signal switch to distribute to the
lamp)
2.1.1. Circuit diagram
To generate the frequency for the turn signal, a turn signal relay is used in the turn signal circuit. The current from the turn signal relay will be sent to the turn signal switch assembly to distribute the current to the turn signal lights for the driver's purpose.
Figure 3.3. Schematic diagram of a turn signal circuit without a hazard switch
1. Battery; 2. Electric lock; 3. Turn signal relay; 4. Turn signal switch; 5. Turn signal lamp; 6. Turn signal lamp; 7. Hazard switch
Figure 3.4 Schematic diagram of turn signal circuit with hazard switch
1. Battery; 2. Combination switch cluster; 3. Turn signal;
4. Turn signal light; 5. Turn signal relay
Today's cars no longer use three-pin turn signal relays (B, L, E) but use eight-pin turn signal relays (figure 3.5) (pin number 8 is used for hazard lights).
For this type, the current supplying the turn signal lights is supplied directly from the turn signal relay to the lights.
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s3 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; vertical-align: 1pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -9pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s9 { color: #008000; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; te -
 Evaluating the effectiveness of agricultural and forestry farming models on limestone terrain in Yen Minh district, Ha Giang province as a basis for planning the district's agricultural and forestry development - 10
Evaluating the effectiveness of agricultural and forestry farming models on limestone terrain in Yen Minh district, Ha Giang province as a basis for planning the district's agricultural and forestry development - 10 -
 Lessons Learned From Related Studies For M&A Activities In Vietnam
Lessons Learned From Related Studies For M&A Activities In Vietnam -
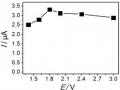
 Effects of Voltage (A); Kcl Concentration (B); Electrolysis Time (C); and Stirring Time (D) on Hg Signal (Ii)
Effects of Voltage (A); Kcl Concentration (B); Electrolysis Time (C); and Stirring Time (D) on Hg Signal (Ii) -
 De Gruyter, C., Currie, G., Rose, G. (2017). Sustainability Measures Of Urban Public Transport In Cities: A World Review And Focus On The Asia/middle East Region . Sustainability, Vol. 9, No. 1,
De Gruyter, C., Currie, G., Rose, G. (2017). Sustainability Measures Of Urban Public Transport In Cities: A World Review And Focus On The Asia/middle East Region . Sustainability, Vol. 9, No. 1,
- Segment 8 (NS gene), is a gene encoding a non-structural protein, with the most stable length in the influenza A virus genome, about 890 bp in size, encoding the synthesis of two proteins, NS1 and NS2 (also known as NEP, nuclear export protein), which play a role in protecting the virus genome. If they are missing, the virus will be deficient (Urphy and Webster, 1996; Sekellick et al., 2000). The toxicity of the virus is related to this non-structural gene, found in the AH N1 9 variant (Tian et al., 2000). In nature, mutations that delete part of the gene are associated with reduced virulence (Zhou et al., 2007). NS1 has a calculated molecular weight of 2.103 Da (in fact,
25.103 Da), responsible for transporting viral messenger RNA from the nucleus to the cytoplasm of infected cells, and acting on the transporting NAs as well as the cleavage and translation processes of the host cell. NEP or NS2, is a gene formed from two gene segments (30 bp and 336 bp) encoding a protein with a calculated molecular mass of about 14.103 Da (actually 12x103 Da), which plays a role in transporting viral NPs out of the nucleus of infected cells to assemble with the capsid to form new virus particles (Sekellick et al., 2000; Zhu et al., 2008).
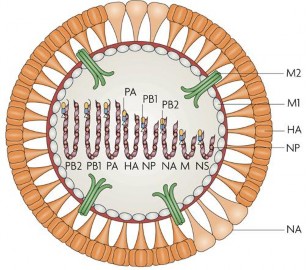
Figure 1.4. Genome model ic A (St bb, 1965
Segments 4 and 6 encode the capsid surface proteins (HA and NA) of the virus, which have antigenic properties specific to each influenza A virus strain. In other words, the HA and NA proteins allow the identification of the subtype of influenza A virus in general and avian influenza virus in particular.
Thus, influenza A virus (type: influenza A/H5N1) has a genome structured from 8 separate segments and does not have a gene encoding the NA repair enzyme, creating favorable conditions for the appearance of point mutations in genome gene segments through the process of virus replication, or the exchange of gene segments between influenza virus strains co-infecting the same cell, which can very likely lead to changes in antigenic properties creating new influenza A virus strains (Suarez and Schultz-Cherry, 2000).
1.3.2. The IC chip is not stable.
Influenza A virus is subtyped based on the surface antigens (proteins) HA (Hemagglutinin - abbreviated as H) and NA (Neuraminidase - abbreviated as N) which play an important role in protective immunity. Hemagglutinin is considered to be the factor that determines both the antigenicity and virulence of influenza A virus.
1.3.2.1. HA Protein (Hemagglutinin)
Hemagglutinin protein is a glycoprotein belonging to type I membrane protein (lectin), which has the ability to cause agglutination of chicken red blood cells in vitro . Specific antibodies to HA can block that agglutination, which is called anti-HA.
There are 16 HA subtypes that have been identified (H1 - H16), the H16 subtype was recently identified in the Swedish albatross virus (1999), three subtypes (H1, H2 and H3) that are adapted to infect humans have been implicated in historical influenza pandemics (Urphy and Webster, 1996). There are approximately 400 HA molecules on the surface of a virus's capsid, which play an important role in virus recognition and initiation of virus infection into host cells (Bender et al., 1999; Wagner et al., 2002). HA molecule has a trigonal shape, about 130 angstroms (Å) long, consisting of 3 monomers, each monomer is made up of two subunits HA1 (36 kDa) and HA2 (2 kDa), linked together by disulfide bridges (-SS-). After synthesis, the monomers have been glycosylated and attached to the outer surface of the capsid as the HA2 subunit, the free head is spherical and shaped under the HA1 unit containing the appropriate HA receptor binding site on the surface of the target cell membrane (Bosch et al., 1981; Wagner et al., 2002).
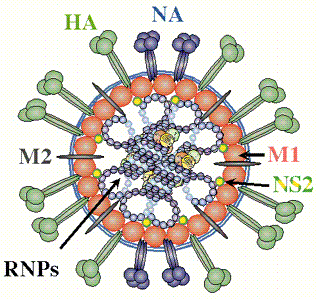
Figure 1.5. Heltin and Neuraminidase inhibitory activity assay (www.aht.org.uk)
The combination of HA with specific receptors (glycoproteins containing sialic acid) on the surface of the cell membrane, at the beginning of the virus infection process on the host, helps the virus penetrate, fuse with the membrane and release the NA genome to carry out the process.
replication process in infected cells. The association process depends on the spatial conformation of the sialic acid receptor of the target cell with the receptor binding site on the HA molecule of the influenza virus, determining the ease of virus invasion of different host species (Wagner et al., 2002; Wasilenko et al., 2008). Amino acid position 226 (aa226) of HA 1 has been identified as the decisive site for HA binding to its specific receptor, most naturally circulating influenza A virus strains have this site as Glycine, which is adapted to the Gal α-2,3 sialic acid receptor (containing sialic acid linked to the hydroxyl group (4-OH) of galactose with an α-2,3 rotation) of respiratory epithelial cells of birds and poultry (natural hosts of influenza A virus). In addition, some other amino acid positions: Glutamine 222, Glycine 224, or the SGVSS and NGQSG structures are also closely related to the ability to adapt to the host cell membrane surface containing sialic acid (Luong and Palese, 1993). In particular, some highly virulent AH Nx; AH Nx strains currently circulating can infect humans, when they have a high load in the respiratory tract (due to direct contact with waste or infected poultry) (Bauer et al., 2006; Hui, 2008; Wasilenko et al., 2008).
The HA protein coding sequence, and the HA protein's splicing sequence composition, as well as the amino acid positions related to the ability to bind to adaptive receptors, are considered molecular markers in the study of HA antigen gene analysis (Luong and Palese, 1993). The HA protein is also an important surface antigen of influenza A virus, stimulating the body to produce a humoral immune response specific to each HA type, and participating in the virus neutralization reaction. It is considered a protein that determines both the antigenicity and virulence of the virus, and is the target of immunological protection to prevent the invasion of the infected body by the virus, the basis for the preparation of current influenza vaccines (Bosch et al., 1988; Horimoto and Kawaoka, 2001; Atrosovich et al., 1999).
1.3.2.2. Protein NA (neuraminidase)
Neurominidase protein, also known as sialidase (international code EC 3.2.1.18), is an enzyme protein that is essentially a glycoprotein attached to the surface of the capsid.
of influenza A virus, antigenically specific to each NA subtype (Baigent et al., 2001; Uiprasertkul et al., 2007). There are 9 subtypes (from N1 to N9) found mainly in avian influenza viruses, two subtypes N1 and N2 found in human influenza viruses associated with historical influenza pandemics (Wasilenko et al., 2008). There are about 100 NA molecules interspersed between HA molecules on the surface of the viral capsid. The NA molecule has a mushroom-shaped convex shape, the free end (containing the active site) consists of 4 nm spherical subunits on the same plane, and the hydrophobic part is attached to the capsid shell (Castrucci and Kawaoka, 1993).
The NA protein acts as an enzyme that breaks the bond between the sialic acid residue of the infected cell membrane and the carbohydrate molecule of the HA protein, releasing the virus particles from the infected cell membrane, accelerating the spread of the virus in the host body, and preventing the assembly of new virus particles on the cell membrane. On the other hand, NA participates in breaking this bond during the " fusion " phase, accelerating the "uncoating" process to release the viral genome into the cytoplasm of the infected cell, helping the virus to multiply faster (Uiprasertkul et al., 200). In addition, NA also breaks glycoside bonds, releasing neuraminic acid to dilute the respiratory epithelial surface mucosa, creating conditions for the virus to quickly access epithelial cells and escape non-specific inhibitors. Along with the role of the HA antigen, all three of the above effects of NA participate in increasing the pathogenicity of influenza A virus in the host body. Therefore, NA is the target of current non-specific antiviral drugs and pharmaceuticals, especially Oseltamivir (trade name Tamiflu), which blocks this enzyme, preventing the release of new virus particles from target cells, protecting the body (Aoki et al., 2000; Castrucci and Kawaoka, 1993).
In addition, NA is also a surface antigen of the virus, participating in stimulating the immune system of the host body, producing specific antibodies to the NA antigen of the infecting virus strains that have the effect of blocking the NA protein (Doherty et al., 2006). Thus, the NA antigen along with the HA antigen of the virus are the main targets of the body's immune defense mechanism.
with influenza A virus, and is the basis for the preparation of current influenza vaccines for humans and poultry, to prevent avian influenza outbreaks and limit transmission to humans (Suarez and Schultz-Cherry, 2000; Wu et al., 2008).
1.3.2.3. Non-stop transformation methods
There are three main ways to change influenza A virus antigens (Wu et al., 2008).
a. Antigenic deviation phenomenon
Antigenic drift is essentially point mutations that occur in viral genome segments. Because influenza A virus is an obligate intracellular parasite, there is no “ proof reading ” mechanism during transcription and replication of the target cell nucleus. The lack of NA repair enzymes leads to NA-dependent replication enzymes being able to “ insert ” ( expansion mutation ), delete or replace ( slippage-deletion mutation ) (Le Thanh Hoa et al., 2006) one or more nucleotides that are not repaired in the new single-stranded NA molecule of the virus (Conenello et al., 2000; Urphy and Webster, 1996). Depending on the location of the mutations in the coding triplet, they can directly change the amino acids in the sequence of the expressed encoded protein, leading to changes in protein properties, or accumulate in the gene segment where the mutation occurs (point mutation). The frequency of point mutations is very high, with 1 nucleotide difference for every 10,000 nucleotides (corresponding to the length of the NA genome of influenza A virus) (Prel et al., 2008; Wagner et al., 2002). Thus, almost every newly produced virus particle contains a point mutation in its genome, and these mutations accumulate over many generations of viruses, giving rise to a new virus subtype with new antigenic properties that may be distorted (Figure 1.6).
This phenomenon often occurs in the NA and HA antigen gene segments, creating new amino acid synthesis codes, or changing the structure leading to changes in the properties of that protein, or having a very high glycosylation ability in the structure of the antigen polypeptide chain, creating a new virus variant that changes its pathogenicity or new antigenic properties (Wasilenko et al., 2008, Acken et al., 2006; Chen et al., 2008).
b. Antigen mixing phenomenon
The phenomenon of antigenic shift (also known as exchange or reassortment) of antigenic genes ( antigenic shift ) is only found in influenza viruses, and very few other avian NA viruses, allowing the virus to have a very high mutation ability. The genome consists of 8 separate gene segments of influenza A virus that are exchanged between 2 different influenza A virus strains when co-infected in a cell, so that the reassortment or exchange of gene segments of the two virus strains can occur during the reassortment of the NA genome, creating different states of the NA genome of new virus particles from the two NA genomes of the original viruses. The result is the creation of a new generation of viruses with combined gene segments, and sometimes helps them to be able to infect new host species or increase their virulence (Figure 1. ) (Hilleman, 2002; acken et al., 2006; Chen et al., 2006).
c. Glycosylation phenomenon
Glycosylation is the attachment of a carbohydrate chain ( oligosaccharide ) to the amino acid Asparagine (N) at certain positions in the polypeptide chain HA or NA, or some other polypeptides of the influenza virus. Usually the oligosaccharide chain is attached at the NXS/T position (N = Asparagine; X = any amino acid, except Proline; ST = Serine or Threonine) (Baigent and cCauley, 2001). These are the positions that are thought to bind to antibodies produced by the body in response to antigen stimulation, to protect the infected body. The phenomenon of antigenic deviation causes point mutations in the coding sequence of Asparagine, creating the premise for glycosylation to occur during the synthesis of HA or NA polypeptide chains, changing the expression of antigenic properties of HA and NA, helping the virus escape the protective immune effects of the host and regulating viral replication (Baigent and McCauley, 2001).
The phenomenon of “ antigenic drift ” and “ glycosylation ” occurs continuously over time, while the phenomenon of “ antigenic mixing ” can occur with all strains of influenza A virus, when co-infecting all different host species in the same cell. This is also the worrying problem of the A/H5N1 influenza virus today, although this virus has not adapted to easily infect humans, but it has the ability to cause disease in humans, and it is very possible that AH N1 recombined ( borrowed ) the HA gene or