The mean was 47.44% higher in single mutants (p = 0.0254) and 41.7% higher in double mutants (p = 0.0472).
However, the raffinose content of the single mutant and double mutant lines was not statistically different (p = 0.24). Furthermore, the verbascose content was reduced by 22.6% (in single mutant seeds) and 42.1% (in double mutant seeds) compared to the control.
Total carbohydrates
Total Raffinose
Figure 3.24. Total soluble carbohydrates and total RFOs measured by HPLC (DT26 is the control seed; D1.1-7-2, D1.1-14-3 are seeds from T2 plants with single GmGOLS03 mutant genes; D1.1-5-4, D1.1-7-1 are seeds from T2 plants with double GmGOLS03 GmGOLS19 mutant genes. Statistical analysis was performed using one-way ANOVA. Different letters indicate significant differences at p < 0.05. Mean ± SD are shown for n = 4)
The total RFOs content, measured as the sum of raffinose, stachyose, and verbascose, was 64.7 mg/g dry weight in the control seeds, which decreased by 30.2% to 45.13 mg/g in the single mutant and by 34.1% to 41.95 mg/g in the double mutant. Meanwhile, no significant changes in glucose or fructose content were found between the control and mutant seeds (p > 0.05).
Figure 3.25. Carbohydrate composition in soybeans
(A). The carbohydrates in large amounts are stachyose and sucrose. (B) the carbohydrates in smaller amounts.
DT26 is the control seed; D1.1-7-2, D1.1-14-3 are seeds from T2 plants with single GmGOLS03 mutant gene; D1.1-5-4, D1.1-7-1 are seeds from T2 plants with double GmGOLS03 GmGOLS19 mutant genes. Statistical analysis was performed using one-way ANOVA. Different letters indicate significant differences at p < 0.05. Mean ± SD are shown for n = 4.
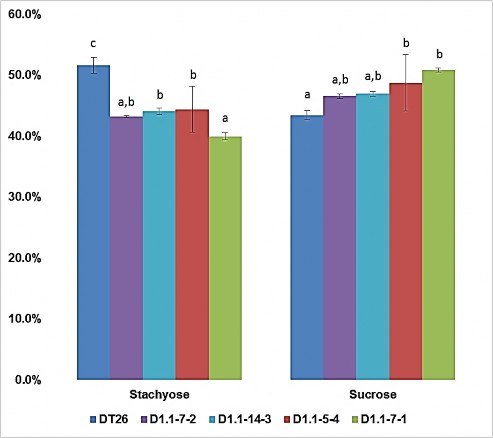
Figure 3.26. The ratio of stachyose and sucrose carbohydrates to the total mass of soluble carbohydrates in soybean seeds
DT26 is the control seed; D1.1-7-2, D1.1-14-3 are seeds from T2 plant with single gene mutation GmGOLS03; D1.1-5-4, D1.1-7-1 are seeds from T2 plant with two genes mutation
GmGOLS03 GmGOLS19. Statistical analysis was performed using one-way ANOVA. Different letters indicate significant differences at p < 0.05. Mean ± SD are shown for n = 4.
Notably, when the individual carbohydrate forms were calculated as a percentage of total soluble carbohydrate (Figure 3.26), it was evident that the relative proportion of sucrose was actually increased in both the single and double mutants compared to the control. In addition, the proportion of stachyose was lower in the single and double mutants than in the control.
3.5.3.2 Analysis of other components in seeds of mutant soybean lines
Quantitative analysis of other grain components including moisture, protein, fat and starch was performed (Figure 3.27, Table 3.5).
Table 3.5 Other components in beans
Protein (%) | Fat (%) | Starch (%) | |
WT (DT26) | 38.30 ± 0.49a | 19.83 ± 1.28a | 5.090 ± 0.84a |
D1.1-7-2 | 40.06 ± 1.09b | 22.88 ± 0.52 b | 5.770 ± 1.00 a |
D1.1-14-3 | 40.06 ± 0.54b | 20.92 ± 0.69a | 4.775 ± 0.38a |
D1.1-5-4 | 38.28 ± 0.49a | 20.37 ± 1.45a | 5.509 ± 1.29a |
D1.1-7-1 | 38.68 ± 0.53a | 20.52 ± 0.79a | 5.219 ± 1.19a |
Maybe you are interested!
-
 Net Interest Income Ratio From Tdtd In The Period 2015-2019
Net Interest Income Ratio From Tdtd In The Period 2015-2019 -
 Classification Table of Overdue Debt Ratio by Collateral of Seabank
Classification Table of Overdue Debt Ratio by Collateral of Seabank -
 Ratio of Management Staff at Vietnam Pharmaceutical Corporation
Ratio of Management Staff at Vietnam Pharmaceutical Corporation -
 Ratio of Income from Wages and Pensions of 15 Households
Ratio of Income from Wages and Pensions of 15 Households -
 Compare Vietnam's Loan-to-Deposit Ratio to Some Other Countries
Compare Vietnam's Loan-to-Deposit Ratio to Some Other Countries
Note: WT (wild type) seeds are the DT26 variety without mutations; D1.1-7-2, D1.1-14-3 are the names of T2 soybean lines with single GmGOLS03 mutant genes; D1.1-5-4, D1.1-7-1 are the names of T2 soybean lines with simultaneous mutations of both GmGOLS03 and GmGOLS19 double mutant genes. The data were statistically analyzed by ANOVA and post hoc tested by Tukey test at a significance level of p=0.05.
Starch Moisture Fat
Protein
Figure 3.27. Proportions of other components in soybeans
DT26: seeds of control line; DT1.1-7-2, DT1.1-14-3: seeds of T2 lines carrying single gene mutation GmGOLS03; DT1.1-5-4, DT1.1-7-1: seeds of T2 lines carrying double gene mutation GmGOLS03 and GmGOLS19
The analysis results showed no significant difference in grain starch composition between the mutant and control seeds tested. However, seeds from the mutant lines
The single mutant GmGOLS03 had increased protein content (40.06% vs. 38.30% of the control, p = 0.00254) and increased fat content (21.9% vs. 19.8% of the control, p = 0.0201), while in the double mutant lines GmGOLS03 and GmGOLS19 there were no significant changes (p = 0.581 and 0.406 for protein and fat, respectively).
These results suggest that the function of GmGOLS03 and GmGOLS19 is not only limited to the biosynthesis of RFOs but may also be involved in the biosynthesis of some other components present in soybean seeds and further studies are needed to clarify this conclusion.
3.6. Testing for off-target mutations and selecting potential mutant lines that do not carry the transgene
To ensure the applicability of promising mutant soybean lines with low raffinose sugar content in seeds, it is important to confirm the absence of off-target mutations as well as to select mutant lines that do not carry the transgene. After selecting soybean lines with stable homozygous mutations, analyze off-target mutations and screen mutant lines that do not carry the transgene.
3.6.1. Off-target mutation analysis
Off-target activity has been reported in several studies using the CRISPR/Cas9 system [83].
Table 3.6 Possible off-target mutations in T2 mutant plants
Sequence | MMs | Gene location | Functional area | Mutations identified | |
Target sequence | GAGTCACACCCCTCAGT ACA | 0 | Glyma.03G38080 Glyma.19G40680 | Exon | |
Off-target may be available | GAGTCA AG CCCC A CAGT ACA TGG | 3 | Glyma.07G34570 | UTR | 0 |
Target sequence | GCACCTTCTCCGGGCATTG C | 0 | Glyma.03G38080 Glyma.19G40680 | Exon | |
Off-target may be available | GCAC T T A CT T CGGG G ATT GC AGG | 4 | Glyma.09G14090 | Exon | 0 |
Note: Red letters in the sequence show the position of deviation from the target sequence, underlined letters are the PAM recognition sequence; MMs: number of positions of deviation from the design orientation sequence.
During target selection using the online program CCTop [93], two potential off-target mutation sites located in the genes Glyma.07G220600 and Glyma.09G169400, respectively , were identified. Both the genome sequences of the control variety William 82 and the test varieties (DT26 and Mr) contained potential off-target sites with identical PAM sequences and 3-4 nucleotide mismatches to any of the targets used (Table 3.6).
We performed off-target analyses on mutant lines T2 and T3 by sequencing off-target regions on two genes Glyma.07G220600 and Glyma.09G169400 (Figure 3.28).
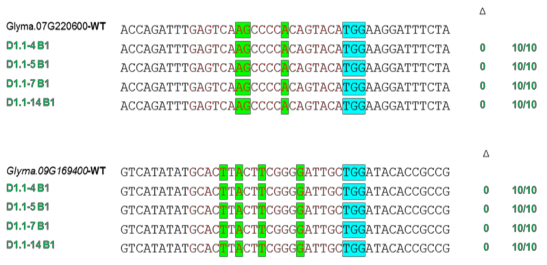
The results showed that no editing activity was detected at off-target sites, indicating high specificity in gene editing using the CRISPR/Cas9 system in soybean.
Figure 3.28. Off-target sequence analysis in the genome of T2 generation mutant soybean lines
The green shaded area shows the sequence that differs from the target sequence, the blue shaded area shows the PAM recognition sequence.
3.6.2. Identification of homozygous mutations that do not carry the transgene
Screening of transferred genes from generation T0 to T2 was carried out by applying herbicide on leaves, combined with PCR analysis of DNA samples to confirm the inheritance of the bar gene . The screening results are shown in Table 3.3 and Figure 3.29. In generation T1, all 29 T1 plant samples of line DT1.1 screened showed herbicide resistance, and also gave positive results when analyzed by PCR with specific primer pairs pcoCas9 and 35SPPDK (Appendix 1), which confirmed the presence of the gene.
In addition, some T1 plants derived from transgenic lines M3.1 and M4.1 gave opposite results, thereby confirming that these T1 plants have genotypes that do not carry the transgene.
Control
Figure 3.29. Identification of mutant lines that do not carry the transgene
Continuing to screen the T2 generation of the DT1.1 line, we identified the DT1.1-7-1 line, with the double GmGOLS mutant genotype , and the DT1.1-7-2 line with a single mutation of the GmGOLS03 gene , both of which did not carry the herbicide resistance gene. All T3 plants generated from these two mutant lines were sensitive to herbicides. PCR analysis also showed that there was no expression of the bar and Cas9 genes in the DNA samples of the T3 plants. This confirmed that the transgenic construct had separated from the mutant lines in the T2 and T3 generations.
From the above analysis results, we have selected potential homozygous mutant lines with low raffinose sugar content in seeds, no transgenes and no off-target mutations. These promising mutant lines are an important source of raw materials for the breeding of Vietnamese soybeans towards improving seed quality in the future.
CHAPTER 4. DISCUSSION
Comparing with previous works in the same field and research direction, we see new discoveries and scientific contributions from the research results in this thesis. We discuss this information in detail in the following sections:
4.1 Efficiency of gene transfer and gene editing of hairy root induction system on Vietnamese soybean varieties
Sustainable and stable gene transfer into soybean plants often uses cotyledons and growing tips as raw materials. However, the efficiency of this method is not high, requires an optimal process and takes a lot of time and cost to complete a process and create transgenic soybean plants. Therefore, the hairy root system is considered an effective method in studies of symbiosis, nutrient absorption, interaction with pathogens and especially in research on gene function or gene editing in soybean.
The results of the hairy root induction system and gene transfer to hairy roots on some soybean varieties studied are presented in section 3.2.1. Compared with the study by Chen and co-authors in 2018, 5-day-old soybean cotyledons including 0.5 cm long petioles were used as transformation materials [81]. The hairy root induction rate of soybean varieties in this study reached 90-99%. However, the gene transfer efficiency was only 30-60%. Similarly, when 5-day-old sprouts were used in gfp gene transformation in the publication by Keyes and co-authors in 2009, the hairy root induction rate was only 45%, of which the number of hairy roots carrying and expressing the transgene was only about 55% [94].
In vitro hairy root induction method with 3-day-old cotyledons of Vietnamese soybean varieties, we obtained the highest gene conversion rate of 79.8% with DT26 variety when using the construct carrying the gfp gene . The hairy root induction rate also reached over 90% with this soybean variety. This method has given gene expression results in a shorter time, helping to reduce the time and cost of testing the activity of the gene transfer constructs.
The CRISPR/Cas9 system has been successfully applied to create genetic mutations.