Table 3.6. Recognition results of Example 2
ID
Decision tree recognition results | |
25 | 1 |
26 | 1 |
27 | 2 |
28 | 2 |
29 | 3 |
30 | 3 |
Maybe you are interested!
-
 Evaluation Results of Cvno Service Quality Criteria and Individual Customer Satisfaction Level for Bidv
Evaluation Results of Cvno Service Quality Criteria and Individual Customer Satisfaction Level for Bidv -
 Identify Rating Levels and Rating Scales
zt2i3t4l5ee
zt2a3gstourism,quan lan,quang ninh,ecology,ecotourism,minh chau,van don,geography,geographical basis,tourism development,science
zt2a3ge
zc2o3n4t5e6n7ts
of the islanders. Therefore, this indicator will be divided into two sub-indicators:
a1. Natural tourism attractiveness a2. Cultural tourism attractiveness
b. Tourist capacity
The two island communes in Quan Lan have different capacities to receive tourists. Minh Chau Commune is home to many standard hotels and resorts, attracting high-income domestic and international tourists. Meanwhile, Quan Lan Commune has many motels mainly built and operated by local people, so the scale and quality are not high, and will be suitable for ordinary tourists such as students.
c. Time of exploitation of Quan Lan Island Commune:
Quan Lan tourism is seasonal due to weather and climate conditions and festivals only take place on certain days of the year, specifically in spring. In Quan Lan commune, the period from April to June and from September to November is considered the best time to visit Quan Lan because the cultural tourism activities are mainly associated with festivals taking place during this time.
Minh Chau island commune:
Tourism exploitation time is all year round, because this is a place with a number of tourist attractions with diverse ecosystems such as Bai Tu Long National Park Research Center, Tram forest, Turtle Laying Beach, so besides coming to the beach for tourism and vacation in the summer, Minh Chau will attract research groups to come for tourism combined with research at other times of the year.
d. Sustainability
The sustainability of ecotourism sites in Quan Lan and Minh Chau communes depends on the sensitivity of the ecosystems to climate changes.
landscape. In general, these tourist destinations have a fairly high level of sustainability, because they are natural ecosystems, planned and protected. However, if a large number of tourists gather at certain times, it can exceed the carrying capacity and affect the sustainability of the environment (polluted beaches, damaged trees, animals moving away from their habitats, etc.), then the sustainability of the above ecosystems (natural ecosystems, human ecosystems) will also be affected and become less sustainable.
e. Location and accessibility
Both island communes have ports to take tourists to visit from Van Don wharf:
- Quan Lan – Van Don traffic route:
Phuc Thinh – Viet Anh high-speed boat and Quang Minh high-speed boat, depart at 8am and 2pm from Van Don to Quan Lan, and at 7am and 1pm from Quan Lan to Van Don. There are also wooden boats departing at 7am and 1pm.
- Van Don - Minh Chau traffic route:
Chung Huong high-speed train, Minh Chau train, morning 7:30 and afternoon 13:30 from Van Don to Minh Chau, morning 6:30 and afternoon 13:00 from Minh Chau to Van Don.
f. Infrastructure
Despite receiving investment attention, the issue of infrastructure and technical facilities for tourism on Quan Lan Island is still an issue that needs to be resolved because it has a direct impact on the implementation of ecotourism activities. The minimum conditions for serving tourists such as accommodation, electricity, water, communication, especially medical services, and security work need to be given top priority. Ecotourism spots in Minh Chau commune are assessed to have better infrastructure and technical facilities for tourism because there are quite complete and synchronous conditions for serving tourists, meeting many needs of domestic and foreign tourists.
3.2.1.4. Determine assessment levels and assessment scales
Corresponding to the levels of each criterion, the index is the score of those levels in the order of 4, 3, 2, 1 decreasing according to the standard of each level: very attractive (4), attractive (3), average (2), less attractive (1).
3.2.1.5. Determining the coefficients of the criteria
For the assessment of DLST in the two communes of Quan Lan and Minh Chau islands, the students added evaluation coefficients to show the importance of the criteria and indicators as follows:
Coefficient 3 with criteria: Attractiveness, Exploitation time. These are the 2 most important criteria for attracting tourists to tourism in general and eco-tourism in particular, so they have the highest coefficient.
Coefficient 2 with criteria: Capacity, Infrastructure, Location and accessibility . Because the assessment area is an island commune of Van Don district, the above criteria are selected by the author with appropriate coefficients at the average level.
Coefficient 1 with criteria: Sustainability. Quan Lan has natural and human-made ecotourism sites, with high biodiversity and little impact from local human factors. Most of the ecotourism sites are still wild, so they are highly sustainable.
3.2.1.6. Results of DLST assessment on Quan Lan island
a. Assessment of the potential for natural tourism development
For Minh Chau commune:
+ Natural tourism attractiveness is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined as average (2 points) and the coefficient is quite important (coefficient 2), then the score of Capacity criterion is 2 x 2 = 4.
+ Exploitation time is long (4 points), the most important coefficient (coefficient 3) so the score of the Exploitation time criterion is 4 x 3 = 12.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is assessed as good (3 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 3 x 2 = 6 points.
The total score for evaluating DLST in Minh Chau commune according to 6 evaluation criteria is determined as: 12 + 4 + 12 + 4 + 4 + 6 = 42 points
Similar assessment for Quan Lan commune, we have the following table:
Table 3.3: Assessment of the potential for natural ecotourism development in Quan Lan and Minh Chau communes
Attractiveness of self-tourismof course
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
CommuneMinh Chau
12
12
4
8
12
12
4
4
4
8
6
8
42/52
Quan CommuneLan
6
12
6
8
9
12
4
4
4
8
4
8
33/52
b. Assessment of the potential for humanistic tourism development
For Quan Lan commune:
+ The attractiveness of human tourism is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined to be large (3 points) and the coefficient is quite important (coefficient 2), then the score of the Capacity criterion is 3 x 2 = 6.
+ Mining time is average (3 points), the most important coefficient (coefficient 3) so the score of the Mining time criterion is 3 x 3 = 9.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points.
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is rated as average (2 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 2 x 2 = 4 points.
The total score for evaluating DLST in Quan Lan commune according to 6 evaluation criteria is determined as: 12 + 6 + 6 + 4 + 4 + 4 = 36 points.
Similar assessment with Minh Chau commune we have the following table:
Table 3.4: Assessment of the potential for developing humanistic eco-tourism in Quan Lan and Minh Chau communes
Attractiveness of human tourismliterature
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Quan CommuneLan
12
12
6
8
9
12
4
4
4
8
4
8
39/52
Minh CommuneChau
6
12
4
8
12
12
4
4
4
8
6
8
36/52
Basically, both Minh Chau and Quan Lan localities have quite favorable conditions for developing ecotourism. However, Quan Lan commune has more advantages to develop ecotourism in a humanistic direction, because this is an area with many famous historical relics such as Quan Lan Communal House, Quan Lan Pagoda, Temple worshiping the hero Tran Khanh Du, ... along with local festivals held annually such as the wind praying ceremony (March 15), Quan Lan festival (June 10-19); due to its location near the port and long exploitation time, the beaches in Quan Lan commune (especially Quan Lan beach) are no longer hygienic and clean to ensure the needs of tourists coming to relax and swim; this is also an area with many beautiful landscapes such as Got Beo wind pass, Ong Phong head, Voi Voi cave, but the ability to access these places is still very limited (dirt hill road, lots of gravel and rocks), especially during rainy and windy times; In addition, other natural resources such as mangrove forests and sea worms have not been really exploited for tourism purposes and ecotourism development. On the contrary, Minh Chau commune has more advantages in developing ecotourism in the direction of natural tourism, this is an area with diverse ecosystems such as at Rua De Beach, Bai Tu Long National Park Conservation Center...; Minh Chau beach is highly appreciated for its natural beauty and cleanliness, ranked in the top ten most beautiful beaches in Vietnam; Minh Chau commune is also home to Tram forest with a large area and a purity of up to 90%, suitable for building bridges through the forest (a very effective type of natural ecotourism currently applied by many countries) for tourists to sightsee, as well as for the purpose of studying and researching.
Figure 3.1: Thenmala Forest Bridge (India) Source: https://www.thenmalaecotourism.com/(August 21, 2019)
3.2.2. Using SWOT matrix to evaluate Quan Lan island tourism
General assessment of current tourism activities of Quan Lan island is shown through the following SWOT matrix:
Table 3.5: SWOT matrix evaluating tourism activities on Quan Lan island
Internal agent
Strengths- There is a lot of potential for tourism development, especially natural ecotourism and humanistic ecotourism.- The unskilled labor force is relatively abundant.- resource environmentunpolluted, still
Weaknesses- Poorly developed infrastructure, especially traffic routes to tourist destinations on the island.- The team of professional staff is still weak.- Tourism products in general
quite wild, originalintact
general and DLST in particularalone is monotonous.
External agents
Opportunity- Tourism is a key industry in the socio-economic development strategy of the province and Van Don economic zone.- Quan Lan was selected as a pilot area for eco-tourism development within the framework of the green growth project between Quang Ninh province and the Japanese organization JICA.- The flow of tourists and especially ecotourism in the world tends toincreasing
Challenge- Weather and climate change abnormally.- Competition in tourism products is increasingly fierce, especially with other localities in the province such as Ha Long, Mong Cai...- Awareness of tourists, especially domestic tourists, about ecotourism and nature conservation is not high.
Through summary analysis using SWOT matrix we see that:
To exploit strengths and take advantage of opportunities, it is necessary to:
- Diversify products and service types (build more tourism routes aimed at specific needs of tourists: experiential tourism immersed in nature, spiritual cultural tourism...)
- Effective exploitation of resources and differentiated products (natural resources and human resources)
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s3 { color: #0D0D0D; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -3pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -2pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -1pt; } div.maincontent .s9 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s11 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s12 { color: black; font-family:Symbol, serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s13 { color: black; font-family:Wingdings; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s14 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s15 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s16 { color: black; font-family:Cambria, serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s17 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s18 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s19 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s20 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 10pt; } div.maincontent .s21 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 11pt; } div.maincontent .s22 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s23 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s24 { color: #212121; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; tex
Identify Rating Levels and Rating Scales
zt2i3t4l5ee
zt2a3gstourism,quan lan,quang ninh,ecology,ecotourism,minh chau,van don,geography,geographical basis,tourism development,science
zt2a3ge
zc2o3n4t5e6n7ts
of the islanders. Therefore, this indicator will be divided into two sub-indicators:
a1. Natural tourism attractiveness a2. Cultural tourism attractiveness
b. Tourist capacity
The two island communes in Quan Lan have different capacities to receive tourists. Minh Chau Commune is home to many standard hotels and resorts, attracting high-income domestic and international tourists. Meanwhile, Quan Lan Commune has many motels mainly built and operated by local people, so the scale and quality are not high, and will be suitable for ordinary tourists such as students.
c. Time of exploitation of Quan Lan Island Commune:
Quan Lan tourism is seasonal due to weather and climate conditions and festivals only take place on certain days of the year, specifically in spring. In Quan Lan commune, the period from April to June and from September to November is considered the best time to visit Quan Lan because the cultural tourism activities are mainly associated with festivals taking place during this time.
Minh Chau island commune:
Tourism exploitation time is all year round, because this is a place with a number of tourist attractions with diverse ecosystems such as Bai Tu Long National Park Research Center, Tram forest, Turtle Laying Beach, so besides coming to the beach for tourism and vacation in the summer, Minh Chau will attract research groups to come for tourism combined with research at other times of the year.
d. Sustainability
The sustainability of ecotourism sites in Quan Lan and Minh Chau communes depends on the sensitivity of the ecosystems to climate changes.
landscape. In general, these tourist destinations have a fairly high level of sustainability, because they are natural ecosystems, planned and protected. However, if a large number of tourists gather at certain times, it can exceed the carrying capacity and affect the sustainability of the environment (polluted beaches, damaged trees, animals moving away from their habitats, etc.), then the sustainability of the above ecosystems (natural ecosystems, human ecosystems) will also be affected and become less sustainable.
e. Location and accessibility
Both island communes have ports to take tourists to visit from Van Don wharf:
- Quan Lan – Van Don traffic route:
Phuc Thinh – Viet Anh high-speed boat and Quang Minh high-speed boat, depart at 8am and 2pm from Van Don to Quan Lan, and at 7am and 1pm from Quan Lan to Van Don. There are also wooden boats departing at 7am and 1pm.
- Van Don - Minh Chau traffic route:
Chung Huong high-speed train, Minh Chau train, morning 7:30 and afternoon 13:30 from Van Don to Minh Chau, morning 6:30 and afternoon 13:00 from Minh Chau to Van Don.
f. Infrastructure
Despite receiving investment attention, the issue of infrastructure and technical facilities for tourism on Quan Lan Island is still an issue that needs to be resolved because it has a direct impact on the implementation of ecotourism activities. The minimum conditions for serving tourists such as accommodation, electricity, water, communication, especially medical services, and security work need to be given top priority. Ecotourism spots in Minh Chau commune are assessed to have better infrastructure and technical facilities for tourism because there are quite complete and synchronous conditions for serving tourists, meeting many needs of domestic and foreign tourists.
3.2.1.4. Determine assessment levels and assessment scales
Corresponding to the levels of each criterion, the index is the score of those levels in the order of 4, 3, 2, 1 decreasing according to the standard of each level: very attractive (4), attractive (3), average (2), less attractive (1).
3.2.1.5. Determining the coefficients of the criteria
For the assessment of DLST in the two communes of Quan Lan and Minh Chau islands, the students added evaluation coefficients to show the importance of the criteria and indicators as follows:
Coefficient 3 with criteria: Attractiveness, Exploitation time. These are the 2 most important criteria for attracting tourists to tourism in general and eco-tourism in particular, so they have the highest coefficient.
Coefficient 2 with criteria: Capacity, Infrastructure, Location and accessibility . Because the assessment area is an island commune of Van Don district, the above criteria are selected by the author with appropriate coefficients at the average level.
Coefficient 1 with criteria: Sustainability. Quan Lan has natural and human-made ecotourism sites, with high biodiversity and little impact from local human factors. Most of the ecotourism sites are still wild, so they are highly sustainable.
3.2.1.6. Results of DLST assessment on Quan Lan island
a. Assessment of the potential for natural tourism development
For Minh Chau commune:
+ Natural tourism attractiveness is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined as average (2 points) and the coefficient is quite important (coefficient 2), then the score of Capacity criterion is 2 x 2 = 4.
+ Exploitation time is long (4 points), the most important coefficient (coefficient 3) so the score of the Exploitation time criterion is 4 x 3 = 12.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is assessed as good (3 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 3 x 2 = 6 points.
The total score for evaluating DLST in Minh Chau commune according to 6 evaluation criteria is determined as: 12 + 4 + 12 + 4 + 4 + 6 = 42 points
Similar assessment for Quan Lan commune, we have the following table:
Table 3.3: Assessment of the potential for natural ecotourism development in Quan Lan and Minh Chau communes
Attractiveness of self-tourismof course
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
CommuneMinh Chau
12
12
4
8
12
12
4
4
4
8
6
8
42/52
Quan CommuneLan
6
12
6
8
9
12
4
4
4
8
4
8
33/52
b. Assessment of the potential for humanistic tourism development
For Quan Lan commune:
+ The attractiveness of human tourism is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined to be large (3 points) and the coefficient is quite important (coefficient 2), then the score of the Capacity criterion is 3 x 2 = 6.
+ Mining time is average (3 points), the most important coefficient (coefficient 3) so the score of the Mining time criterion is 3 x 3 = 9.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points.
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is rated as average (2 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 2 x 2 = 4 points.
The total score for evaluating DLST in Quan Lan commune according to 6 evaluation criteria is determined as: 12 + 6 + 6 + 4 + 4 + 4 = 36 points.
Similar assessment with Minh Chau commune we have the following table:
Table 3.4: Assessment of the potential for developing humanistic eco-tourism in Quan Lan and Minh Chau communes
Attractiveness of human tourismliterature
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Quan CommuneLan
12
12
6
8
9
12
4
4
4
8
4
8
39/52
Minh CommuneChau
6
12
4
8
12
12
4
4
4
8
6
8
36/52
Basically, both Minh Chau and Quan Lan localities have quite favorable conditions for developing ecotourism. However, Quan Lan commune has more advantages to develop ecotourism in a humanistic direction, because this is an area with many famous historical relics such as Quan Lan Communal House, Quan Lan Pagoda, Temple worshiping the hero Tran Khanh Du, ... along with local festivals held annually such as the wind praying ceremony (March 15), Quan Lan festival (June 10-19); due to its location near the port and long exploitation time, the beaches in Quan Lan commune (especially Quan Lan beach) are no longer hygienic and clean to ensure the needs of tourists coming to relax and swim; this is also an area with many beautiful landscapes such as Got Beo wind pass, Ong Phong head, Voi Voi cave, but the ability to access these places is still very limited (dirt hill road, lots of gravel and rocks), especially during rainy and windy times; In addition, other natural resources such as mangrove forests and sea worms have not been really exploited for tourism purposes and ecotourism development. On the contrary, Minh Chau commune has more advantages in developing ecotourism in the direction of natural tourism, this is an area with diverse ecosystems such as at Rua De Beach, Bai Tu Long National Park Conservation Center...; Minh Chau beach is highly appreciated for its natural beauty and cleanliness, ranked in the top ten most beautiful beaches in Vietnam; Minh Chau commune is also home to Tram forest with a large area and a purity of up to 90%, suitable for building bridges through the forest (a very effective type of natural ecotourism currently applied by many countries) for tourists to sightsee, as well as for the purpose of studying and researching.
Figure 3.1: Thenmala Forest Bridge (India) Source: https://www.thenmalaecotourism.com/(August 21, 2019)
3.2.2. Using SWOT matrix to evaluate Quan Lan island tourism
General assessment of current tourism activities of Quan Lan island is shown through the following SWOT matrix:
Table 3.5: SWOT matrix evaluating tourism activities on Quan Lan island
Internal agent
Strengths- There is a lot of potential for tourism development, especially natural ecotourism and humanistic ecotourism.- The unskilled labor force is relatively abundant.- resource environmentunpolluted, still
Weaknesses- Poorly developed infrastructure, especially traffic routes to tourist destinations on the island.- The team of professional staff is still weak.- Tourism products in general
quite wild, originalintact
general and DLST in particularalone is monotonous.
External agents
Opportunity- Tourism is a key industry in the socio-economic development strategy of the province and Van Don economic zone.- Quan Lan was selected as a pilot area for eco-tourism development within the framework of the green growth project between Quang Ninh province and the Japanese organization JICA.- The flow of tourists and especially ecotourism in the world tends toincreasing
Challenge- Weather and climate change abnormally.- Competition in tourism products is increasingly fierce, especially with other localities in the province such as Ha Long, Mong Cai...- Awareness of tourists, especially domestic tourists, about ecotourism and nature conservation is not high.
Through summary analysis using SWOT matrix we see that:
To exploit strengths and take advantage of opportunities, it is necessary to:
- Diversify products and service types (build more tourism routes aimed at specific needs of tourists: experiential tourism immersed in nature, spiritual cultural tourism...)
- Effective exploitation of resources and differentiated products (natural resources and human resources)
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s3 { color: #0D0D0D; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -3pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -2pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -1pt; } div.maincontent .s9 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s11 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s12 { color: black; font-family:Symbol, serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s13 { color: black; font-family:Wingdings; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s14 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s15 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s16 { color: black; font-family:Cambria, serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s17 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s18 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s19 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s20 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 10pt; } div.maincontent .s21 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 11pt; } div.maincontent .s22 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s23 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s24 { color: #212121; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; tex -
 Evaluation Results of Quality Indicators and Importance of Each Indicator
Evaluation Results of Quality Indicators and Importance of Each Indicator -
 Evaluation of human resource training quality at ÊMM Hue Hotel - 17
Evaluation of human resource training quality at ÊMM Hue Hotel - 17 -
 Evaluation of the effect of DH fertilizer on growth, yield and quality of some fruit trees at Thai Nguyen University of Agriculture and Forestry - 2
Evaluation of the effect of DH fertilizer on growth, yield and quality of some fruit trees at Thai Nguyen University of Agriculture and Forestry - 2
3.4. Conclusion of chapter 3
In the first part, the author briefly presented the structure of the networks and learning methods of four classic single recognition models, which are multi-layer feedforward neural network MLP, fuzzy logic neural network TSK, support vector machine SVM and random forest RF.
In the next part, the author presents a single recognition model using popular neural networks to recognize electrocardiogram signals such as MLP, TSK, SVM and RF neural networks and builds a coordinated recognition model of the above neural networks to increase the accuracy of the recognition problem.
The next chapter will present the computational results of the proposed recognition methods in chapter 3.
Chapter 4. CALCULATION AND SIMULATION RESULTS
Chapter 4 presents the method of constructing a sample dataset from the MIT-BIH and MGH/MF databases; with this sample dataset, the author uses single neural network models and combined neural network models to identify ECG electrocardiogram signals by simulating on the computer, then evaluates the results to demonstrate the proposed solution of the thesis.
4.1. Building sample data sets
4.1.1. MIT-BIH Database
In this thesis, we use sample ECG signals taken from the famous MIT-BIH arrhythmia database [15] (which can be downloaded from www.physionet.org). There have been many works on ECG signal classification using this database to build and test recognition models, such as in studies [5, 20, 25, 26, 27, 28, 29, 32].
For the purpose of comparison with previous works, this thesis will use the same data set as in studies [5, 28], specifically the identification of arrhythmias originating from the basis set of QRS segments of electrocardiogram signals from 19 patients (database codes are 100, 105, 106, 109, 111, 114, 116, 118, 119, 124,
200, 202, 207, 208, 209, 212, 214, 221 and 222). Use lead I recording for all selected recordings. The selected patients often had more than one arrhythmia in their recordings, the worst case being patient 207 having all 7 types of arrhythmias, details are presented in the appendix.
In the main test of the thesis, 7 types of arrhythmias were considered: Left bundle branch block (L), Right bundle branch block (R ), Atrial premature beat (A), Ventricular premature beat ( V), Ventricular flutter wave (I) , Ventricular escape beat (E) and normal sinus rhythm (N). Because the disease samples were not evenly distributed in the records, most of the patients had normal rhythm N, diseased rhythms appeared very rarely such as ventricular fibrillation I, ventricular escape beat E. To build data sets with diseased patterns that could appear in the same patient (as in the case of patient 207, all six types of diseased rhythms appeared , so in the thesis, all rhythms of all records will not be taken as a database. In the records of the 19 patients above, 6678 samples were taken and divided into two data sets: 3611 samples. for the learning process (to build the model) and 3068 samples used for testing purposes (to check the reliability). The detailed number of samples taken from the records of 19 patients is detailed in Table 4.1.
Table 4.1. Distribution of the number of training samples and test samples of 7 types of arrhythmias from the MIT-BIH database
Rhythm type
Total number of samples | Number of samples | Number of test samples | |
N | 2000 | 1065 | 935 |
L | 1200 | 639 | 561 |
R | 1000 | 515 | 485 |
A | 902 | 504 | 398 |
V | 964 | 549 | 451 |
I | 472 | 271 | 201 |
E | 105 | 68 | 37 |
Total | 6643 | 3611 | 3068 |
Table 4.2. Table dividing the number of study samples and test samples of the 2 types of rhythms
Rhythm type
Total number of samples | Number of samples | Number of test samples | |
Normal | 2000 | 1065 | 935 |
Abnormal | 4643 | 2546 | 2133 |
Total | 6643 | 3611 | 3068 |
In Table 4.1 is the number of heart rate types used in the learning and testing of the recognition model. The limited number of heart rate types (e.g. E, I beats) is due to the limitation in the MIT-BIH database [15]. The original number of normal heart rates N is very high, but we consider these as simple beats for classification and to make the final results generally more independent of the number of samples in each group, the author limits the number of heart rates used in the experiments to a reasonable level usually around 1000 beats.
To generate the feature vector and corresponding output signal from the ECG signal records in the MIT-BIH database, we follow the following procedure:
- QRS complex separation: Because in the MIT-BIH database, ECG signals have been marked with R peak positions and have been labeled with disease classification by specialized doctors, so for each QRS complex sample, in this case, there is no need to use the R peak determination algorithm as presented in section 2.2, but only need to sequentially read the R peak positions of the QRS complex in the ECG signal line, separate the QRS complex by cutting a 250ms window around the R peak (125ms before and 125ms after the R peak position).
1
- Expand the QRS complex extracted above according to the Hermite polynomials according to formula 4.1 to determine the first 16 expansion coefficients as characteristics.
n ( x )
2 n n ! 2e
x 2
2 H n ( x )
(4.1)
- Determine the RR distance from the considered R peak to the previous R peak to make the 17th feature. The average value of the last 10 RR segments will be the 18th feature of the considered QRS complex.
Thus, each QRS complex is separated into 18 characteristics, the output is the code of the disease type of the rhythm under consideration (already marked in the MIT-BIH database ), for example with 7 different rhythm types the corresponding output will be 7 channels with values 0 and 1 (6 channels are 0 and the channel with the code corresponding to the disease type will have value 1).
The following is a superimposed image of the samples of 6 types of disease rhythms with the number as shown in Table 4.1. From Figure 4.1, it can be seen that the major obstacle in recognizing ECG signals is the huge variation in amplitude and shape of heart beats belonging to the same disease type [23]. Moreover, some types of rhythms belonging to different diseases can also be similar in morphology . Therefore, we can see that the problem of classifying ECG signals is a difficult problem.
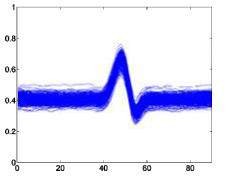
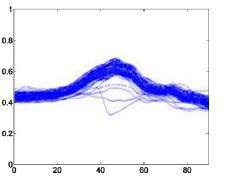
Atrial premature beats – A Ventricular premature beats – E
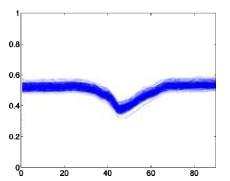
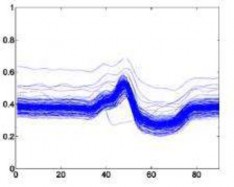
Left bundle branch block –L Right bundle branch block –R
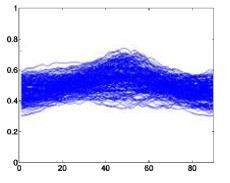
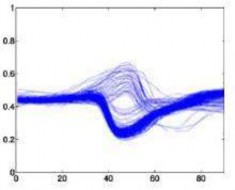
Ventricular Fibrillation – I Ventricular Premature Contractions – V
Figure 4.1. QRS complex patterns of rhythms A, E, L, R, I and V
4.1.2. MGH/MF Database
In addition, the thesis also uses the second database set, MGH/MF [35, 36], this database set includes 250 records of ECG signals, collected from 250 cardiovascular patients in intensive care rooms, operating rooms, cardiac catheterization laboratories, etc. at Massachusetts General Hospital. The thesis chooses to use ECG signal samples of 20 records with code numbers: 029, 030, 058, 105, 106, 107, 108, 110, 111, 114, 117,
119, 121, 123, 124, 125, 128, 131, 137, 142, taking out a total of 4500 samples with 3 types of rhythm: Normal (N -Normal sinus rhythm), premature ventricular contraction (V-Premature ventricular contraction) and supraventricular premature beat (S-Supraventricular premature beat). The detailed number of samples used is listed in Table 4.3 and Table 4.4 below:
Table 4.3. Table of division of number of study samples and test samples of 3 types of rhythms
Rhythm type
Total number of samples | Number of samples | Number of test samples | |
N | 3000 | 1997 | 1003 |
S | 750 | 502 | 248 |
V | 750 | 501 | 249 |
Total | 4500 | 3000 | 1500 |
Table 4.4. Table dividing the number of study samples and test samples of the 2 types of rhythms
Rhythm type
Total number of samples | Number of samples | Number of test samples | |
Normal (normal) | 3000 | 1997 | 1003 |
Abnormal (abnormal) | 1500 | 1003 | 497 |
Total | 4500 | 3000 | 1500 |
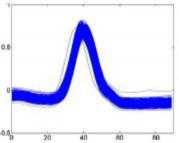
Ventricular extrasystole – V Supraventricular arrhythmia – S Normal – N
Figure 4.2. QRS complex pattern of V, S, N rhythms
To generate the feature vectors and corresponding output signals from the ECG signal records in the MGH/MF database, we proceed similarly to the MIT-BIH database.
4.2. Criteria for evaluating the quality of the ECG signal recognition model
The recognition models (single and combined models) are evaluated using the following criteria:
- Number of incorrectly identified samples;
- FN ( False Negative ): Number of false negative diagnoses, that is, cases where the patient has the disease but is misdiagnosed as normal;
- TN (True Negative ): Number of correct negative diagnoses;
- FP (False Positive): Number of false positive diagnoses, meaning cases where the patient is normal but diagnosed with the disease;
- TP (True Positive ): Number of cases with correct positive diagnosis;
- Sensitivity (True Positive Rate) : The rate of correct positive diagnosis,
Sensitivity
TP TP FN
.100%
(4.2)
- Specificity (True Negative Rate): Rate of correct negative diagnosis
Specificity
TN TN FP
.100%
(4.3)
The recognition model has high quality and reliability when:
- Number of falsely identified samples, number of false positive diagnoses FP, number of false negative diagnoses FN low;
- Correct positive diagnosis rate Sensitivity , correct negative diagnosis rate
The higher the specificity ;
The actual requirements for the quality of ECG signal recognition models are very high. In the criteria for evaluating the model quality listed above, pay attention to the FN parameter (number of false negative diagnoses). In practice, if FN is high, it will cause
dangerous for people because the recognition model will not detect the disease when in fact there is a disease, so if the recognition model has a lower FN, the quality of the model is evaluated to increase.
4.3. Single recognition model results
4.3.1. On the MIT-BIH database
a) Test on the dataset in table 4.1
7 types of beat recognition, MLP, SVM, TSK and RF base recognition models
trained independently on the same training dataset , with the following results:
- MLP Network
The structure is as in [6], 18 input neurons (corresponding to 18 values of the feature vector), 20 hidden neurons, there are 7 output neurons (corresponding to 7 types of arrhythmia),
- TSK Network
The selection structure is as in [5, 6], with 21 inference rules and 7 outputs;
- Support Vector Machine SVM
The selection structure is like [31], the SVM model classifies 7 classes, using the method
OVO should have 21 binary SVM components;
- RF random forest
Following the method of L. Breiman (2001) [19], the RF random forest has 100 component decision trees.
The test results of the basic recognition models on the same test data set in table 4.1, the results are presented in matrix form as in tables 4.5 to 4.9: In which the columns are the standard results (pre-marked), the rows are the recognition results, the main diagonal of the matrix is the number of correctly recognized samples, the values outside the main diagonal are the incorrectly recognized samples, the specific results are as follows:
Table 4.5. Distribution matrix of recognition results of 7 types of rhythm patterns using MLP network
Sample
Result
N | L | R | A | V | I | E | |
N | 905 | 5 | 4 | 5 | 4 | 0 | 0 |
L | 5 | 536 | 0 | 0 | 2 | 0 | 1 |
R | 3 | 0 | 476 | 5 | 1 | 0 | 1 |
A | 17 | 6 | 2 | 382 | 6 | 4 | 0 |
V | 3 | 9 | 1 | 5 | 429 | 2 | 0 |
I | 2 | 3 | 2 | 0 | 7 | 195 | 0 |
E | 0 | 2 | 0 | 1 | 2 | 0 | 35 |
Total error | 30 | 25 | 9 | 16 | 22 | 6 | 2 |
Table 4.6. Distribution matrix of recognition results of 7 types of rhythm patterns using TSK network
Sample
Result
N | L | R | A | V | I | E | |
N | 920 | 8 | 2 | 9 | 4 | 0 | 0 |
L | 0 | 518 | 0 | 1 | 3 | 0 | 0 |
R | 4 | 0 | 481 | 12 | 1 | 0 | 0 |
A | 9 | 19 | 0 | 375 | 2 | 2 | 0 |
V | 2 | 14 | 1 | 1 | 439 | 0 | 0 |
I | 0 | 0 | 1 | 0 | 3 | 199 | 0 |
E | 0 | 2 | 0 | 0 | 0 | 0 | 37 |
Total error | 15 | 43 | 4 | 23 | 13 | 2 | 0 |
Table 4.7. Distribution matrix of recognition results of 7 types of rhythm patterns using SVM network
Sample
Result
N | L | R | A | V | I | E | |
N | 919 | 7 | 1 | 7 | 0 | 0 | 0 |
L | 2 | 546 | 1 | 1 | 3 | 3 | 1 |
R | 1 | 0 | 479 | 4 | 0 | 0 | 0 |
A | 11 | 2 | 2 | 385 | 0 | 0 | 0 |
V | 2 | 5 | 1 | 1 | 448 | 1 | 0 |
I | 0 | 1 | 1 | 0 | 0 | 197 | 0 |
E | 0 | 0 | 0 | 0 | 0 | 0 | 36 |
Total error | 16 | 15 | 6 | 13 | 3 | 4 | 1 |
Table 4.8. Distribution matrix of the results of identifying 7 types of rhythm patterns using RF
Sample Results
N | L | R | A | V | I | E | |
N | 914 | 6 | 3 | 11 | 0 | 0 | 0 |
L | 1 | 547 | 0 | 0 | 4 | 0 | 1 |
R | 1 | 0 | 478 | 4 | 0 | 0 | 0 |
A | 18 | 4 | 1 | 382 | 3 | 0 | 0 |
V | 1 | 3 | 1 | 1 | 443 | 3 | 0 |
I | 0 | 1 | 2 | 0 | 1 | 198 | 0 |
E | 0 | 0 | 0 | 0 | 0 | 0 | 36 |





