and the other copy remains at the old location. In the conservative pathway (as in the case of Tn10) there is no replication. Instead, the element is cut out of the chromosome or plasmid and inserted into the new location. This pathway is also known as the "cut and paste" pathway
2. Mobile genetic elements in viruses
Retroviruses are single-stranded RNA viruses that replicate via double-stranded DNA. RNA is copied into DNA by the enzyme reverse transcriptase. The double-stranded DNA is incorporated into the host cell chromosome, from which it is transcribed to produce viral RNA and proteins to form new virus particles. Some retroviruses, such as mouse mammary tumor virus (MMTV) and Rous sarcoma virus (RSV), infect and stimulate cancer tumors. Once incorporated into the chromosome, the double-stranded DNA copy of the viral genome is called a provirus.
Maybe you are interested!
-
 Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 22
Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 22 -
 Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 20
Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 20 -
 Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 16
Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 16 -
 Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 25
Study on species diversity and genetic relationships of the family Gekkonidae in some limestone mountain areas in the Lao People's Democratic Republic - 25 -
 Applied microbial genetics - 1
Applied microbial genetics - 1
The method of reverse transcription of mobile elements is similar to that of retroviruses. The RNA-mediated movement by reverse transcritase to generate cDNA and the insertion of the cDNA into a new position is called retrotransposition.
Retrotransposition creates a copy of the element at a new location, while the original donor molecule remains unchanged. Retrotransposition thus causes minor disruptions and rearrangements of the host cell genome. The genomic changes associated with retrotransposition lead to the silencing or activation of genes, some of which can cause cancer.
3. Mobile genetic elements in fungi
Gery Fink and colleagues were the first to use yeast to study gene regulation in eukaryotes. They isolated thousands of mutants in the HIS4 gene, which encodes an enzyme involved in histidine synthesis. Of the more than 1,500 random HIS4 mutants found, two had an unstable phenotype. These unstable mutants reverted to the wild-type form 1,000 times more frequently than other HIS4 mutants. These mutants introduced a large DNA insertion into the HIS4 gene, which was mediated by one of the yeast Ty factors. There are 35 copies of the Ty1 insertion factor in the yeast genome.
Cloning of these elements from mutant alleles showed that the insertions were not similar to bacterial IS elements or transposons. Instead, they had the characteristics of retroviruses (animal viruses). There was similarity in the structure and gene content of retroviruses and Ty1 elements isolated from HIS4 mutants. Like retroviruses, yeast transposons have long terminal repeat sequences (LTRs), containing thousands of
The 100 base pairs, called the sequence , are located on both sides of the coding segment, both of which contain the gag gene and the pol gene . Retroviruses have at least three genes that encode three proteins involved in replication: the gag gene encodes a protein that denatures the RNA genome. The pol gene encodes the enzyme reverse transcriptase. The env gene encodes the envelope protein. The Ty element contains only the gag and pol genes , but does not contain the env gene (Figure 4.10).
5 bp direct repeat of target DNA
Ty1 clone inserted
Other chromosome transcription
Code translationReverse transcriptase
Reverse transcription
Figure 4.10 Transposition by retrotransposition
Model of retrotransposon-mediated transposition. An RNA transcript from a retrotransposon is converted to DNA by the enzyme reverse transcriptase encoded by the retrotransposon. The DNA copy is inserted into a new location in the genome.
In 1985, J. Bocke and G. Fink demonstrated that the Ty1 element, like a retrovirus, carries out RNA-mediated transfer. They began by modifying a yeast Ty1 element cloned on a plasmid. First, at one end of the element, a promoter is inserted that is activated by the addition of galactose to the medium. Second, an intron from another yeast gene is inserted into the coding region of the Ty transposon. The addition of galactose increases the frequency of transposition of the modified Ty element. This increases the amount of RNA, because galactose stimulates the transcription of Ty RNA starting from the galactose-sensitive promoter.
Questions and Exercises
1. Why are most mutations affecting structural genes recessive compared to wild-type alleles?
2. How does frameshift mutation formation occur?
3. What are the characteristics of mutagens?
4. Explain the mutagenic basis of the following mutagens: 5-bromuracil, nitrous acid and acridine.
5. Describe a type of random failure that leads to mutation.
6. Analyze the similarities and differences between transposition types.
7. What role do inverted sequences play in transposition?
8. Given a nucleotide sequence on mRNA as follows: Wild type: ... 5' AAUCCUUACGGA 3' ... Mutant type: .... 5' AAUCCUACGGA 3' ...
Please tell me what type of mutation causes the above error?
9. Mutations occur in the nucleotide sequence due to mispairing as follows: 5' AGCTGCCTT 3'
3' ACG ATG GAA 5' (template strand)
Which amino acid is involved in the codon whose nucleotide is changed as above?
References
1. Pham Thanh Ho. 2000. Genetics. Education Publishing House.
2. Le Dinh Luong, Phan Cu Nhan (1998). Genetic basis. Education Publishing House.
3. Hoang Trong Phan. 1995. Molecular genetics. Center for Distance Education, Hue University
4. Anthony J.F. Griffiths, Susan R. Wessler, Richard C. Lewontin, William M. Gelbart, David T. Suzuki, Jeffrey H. Miller. 2004. An introduction to genetics analysis. WH Freeman Publishers.
5. Harlt DL, Jones EW 1998. Genetics - Principle and analysis. Jones and Bartlett Publishers, Toronto, Canada.
6. Stansfield WD 1991. Schaum's outline of theory and problems of genetics. McGraw-Hill, Companies, Inc., United States of America.
7. Watson DJ, Baker TA, Bell SP, Gann A., Levine M., Losick R. 2004. Molecular biology of the gene. Benjamine Cummings, San Francisco, United States of America.
Chapter 5
Virus Genetics
I. Characteristics of viruses
1. Diversity in structure and genetic composition
Viruses have very diverse genomes. The genetic machinery of viruses can be double-stranded DNA (dsDNA), single-stranded DNA (ssDNA), double-stranded RNA (dsRNA) or single-stranded RNA (ssRNA). The RNA genome of a virus is a molecule or a segment, single-stranded (+) or (-) polarity, can be circular or linear. The smallest virus has at most 4 genes, the largest virus has several hundred genes. The genome of viruses has diverse structures but all ensure the common requirement of being able to replicate in the host cell to create both the genome for assembling the next generation of virions and the mRNAs to synthesize viral proteins.
2. Host specificity
Each type of virus can infect and parasitize only a limited range of cells called its host range. Viruses recognize host cells using a “lock and key” principle where the proteins on the outside of the virion fit into specific receptor sites on the cell surface. Some viruses have a wide enough host range to infect several species. For example, rabies viruses can infect many mammals including rodents, dogs, and humans. The range can be very narrow, such as many phages that only infect E. coli bacteria .
II. Genetics of bacteriophages (Bacteriophage or phage)
1. Lysis spot formation and phage mutants
Phages are easily detected because during the lytic cycle, a phage-infected cell ruptures and releases phage particles into the medium (Figure 5.1) . The formation of spots is observed.
A large number of bacterial cells (about 10 8 cells) are spread on a solid medium. After a period of growth, a layer of opaque white bacterial cells is formed. If phage is present at the time the bacteria are spread on the medium, it will infect the bacterial cells. The phage-infected cells then lyse and release many new phages. The next generation of phages infects nearby bacteria, and participates in another lysing cycle, these bacteria rupture and release more phages, which can infect other bacteria in the vicinity. The phage infection cycle continues and after several hours, the phage destroys all the bacterial cells of a cell.
transparent plage area distinct from white bacterial cell layer
opaque.
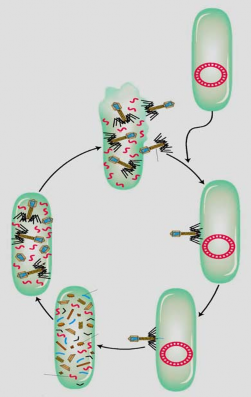
Non-infected cells
Host cell degradation
Phage assembly inside the cell
Free phage
Life cycle
Phage adsorption onto host cells
Phage nucleic acid host cell
The host cell's genetic material is destroyed.
Phage proteins
Phage nucleic acid enters the cell
Phage proteins are synthesized, nucleic acids are replicated, and host cell genetic material is destroyed.
Figure 5.1 Lytic cycle of bacteriophage
Phages can only multiply when grown within bacterial cells, thus depleting the growth medium of nutrients, limiting phage multiplication and spot size. Since each spot is the result of infection with a single initial phage particle, the number of individual spots present on the medium can be counted (Figure 5.2).
The genotypes of phage mutants can be determined by studying the spots. In some cases, the presence of spots is sufficient. For example, phage mutations that reduce the number of progeny phages from infected cells often produce smaller spots. Large spots may be produced by mutations that cause early lysis of infected cells, so that each spot continues to infect more rapidly. Another type of phage mutation is
can be determined by the ability or inability of phages to form spots on particular bacterial strains.
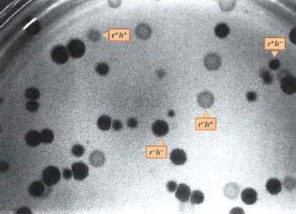
Figure 5.2 Phage entry into host cells in both parental forms simultaneously.
r + : small spot, r - : large spot, h + : faint spot, h - : clear spot
2. Genetic recombination in the lytic cycle
Phages are small in size and must be seen under an electron microscope. But the characteristics of phages are observed based on the spots or host amplitude. Let two phage T 4 strains with different genotypes infect an E. coli bacteria cell , some of the next generation phages will perform genetic recombination, phage DNA replicates and exchanges segments if there is negative interference. Allele r - dissolves quickly, resulting in large spots, allele h - infects host cells, resulting in clear spots. The cross is as follows:
r - h + r + h -
Four types of spots are obtained. The two large, opaque spots and the small, clear spots correspond to the phenotypes of the parental phage. The other two phenotypes, large clear spots and small opaque spots, are recombinant forms corresponding to the genotypes r - h - and r + h + . When many bacteria are infected, a number of reversible recombinant forms are usually found among the phages in the next generation. In the experiment, each of the four genotypes produces a different phenotype in terms of spot type (Figure 5.2) . The number of genotypes can be determined by examining the spots formed. The recombinant frequency, expressed as a percentage, is determined as follows:
Recombination frequency =
ü 1 0 p 0 Täøsänupghage
3. Arrangement of genes in phage chromosomes
Recombination frequencies can be used to determine the distance of a map in eukaryotes. Mapping experiments showed that mutations in T4 mapped into three distinct clusters. All three clusters were linked to one another. George Streisinger et al. (1964) demonstrated that the genetic map of phage T4 was circular.
Membrane
Tail thread
Nucleotide exchange
DNA synthesis, replication and modification
Linkage group of genes determines origin
Tail root disc
Head, neck, neck folds
Tail root disc
Head
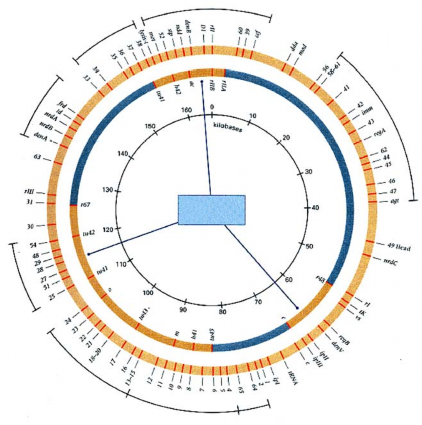
Figure 5.3 Genetic map of T4 with markers
In each cross, three to four genetic markers were set up in turn for each group and run through the entire T4 genome. Several other genes were identified and mapped completely on the circular molecule (Figure 5.3) . The regions in the inner circle are the three clusters of T4 markers that have been identified and mapped. The outer circle contains several large sets of markers that complete the entire circle of the genetic map. The T4 phage genetic map shows that the T4 phage genes cluster extensively according to their function. For example, there is a large cluster of genes for DNA replication in the fourth position.
on the upper right and has the gene cluster that synthesizes the components that make up the phage head at the bottom of the circle.
The T4 phage DNA molecule is a linear single-stranded molecule, with each end of the T4 phage DNA being either replicated or terminally redundant. Thus, each DNA molecule increases in size by 2%. As the DNA is replicated in the cell, recombination between the terminal portions of the T4 genome and homologous sequences of other T4 genomes results in a DNA product that is larger than the head can contain. The duplicated molecules are formed because recombination in the T4 phage genome occurs frequently, with an average of about 20% of recombination events occurring on a chromosome. When the DNA molecule is packaged into the head, it is enzymatically cleaved to contain only about 102% of the length of the T4 phage genome, because of the head repeat.
4. Mapping the fine structure of the rII region of phage T4
Detailed studies of rII mutants of phage T4 shed more light on the gene structure. Wild-type phage T4 r + is capable of simultaneously infecting both E.coli strains B and K. rII mutants infect only strain B but not strain K. Seymour Benzer (1955) obtained 2400 independently derived rII mutants. He crossed the mutants with each other and based on the appearance of wild-type r + recombinant forms , mapped the mutation points.
Each mutant can recombine with other mutants. Deletion mutations prevent recombination with two or more point mutations at different locations in the gene. Each deletion causes the loss of a portion of the phage genome including the rII region. Deletion mapping is a simple method for mapping thousands of mutants. Deletion mapping is based on the presence or absence of recombinant forms. In any cross between an unknown point mutant and a deletion mutant, the appearance of the wild-type form indicates that the point mutation is outside the deletion region. Conversely, if the point mutation occurs within the deletion region, no wild-type recombinant forms will appear in the next generation.
Several crosses were performed to map the detailed mutation of the rII gene. The distances from A1 to A6 and B are shown in Figure 5.4 . A particular mutation was examined located in the A4 region. This mutant did not recombine wild-type in crosses with large deletion mutants such as r1272, r1241, rJ3 and rPT1 but it could recombine wild-type in crosses with rPB242, rA105 and r638. Mutants generated by the same template, resulting from crosses with large deletion mutants, will be ranked





