The sequence of this gene is suitable as a popular model for studying bacterial evolution and taxonomy. The rDNA gene is found in most life forms (except viruses and prions). Components of the rDNA sequences from closely related organisms are markedly similar. This means that the sequences of closely related organisms are precisely aligned, making differences easy to assess. It also means that only a few sets of PCR primers are needed to amplify the rDNA gene from any bacterial species. The 16S rDNA region sequences have been determined for many species. In fact, no other gene is specific to so many species. The length and sequence of the ITS regions of rDNA are thought to be the most rapidly evolving regions and can therefore be highly variable. Universal primers designed from the conserved regions at both ends of the ITS region and the small ITS region (600 ÷ 700 bp) are easily amplified because of the large copy number (up to 30,000 copies per cell) (Dubouzet and Shinoda, 1999) of the rDNA repeat region. This makes the ITS region a subject of interest for evolutionary and phylogenetic studies as well as biogeographic studies (Baldwin et al., 1995) (Figure 1.7).

Figure 1.7. ITS region sequence designed by primer J-RXc 2 of X. axonopodis pv .
Maybe you are interested!
-
 Identify Rating Levels and Rating Scales
zt2i3t4l5ee
zt2a3gstourism,quan lan,quang ninh,ecology,ecotourism,minh chau,van don,geography,geographical basis,tourism development,science
zt2a3ge
zc2o3n4t5e6n7ts
of the islanders. Therefore, this indicator will be divided into two sub-indicators:
a1. Natural tourism attractiveness a2. Cultural tourism attractiveness
b. Tourist capacity
The two island communes in Quan Lan have different capacities to receive tourists. Minh Chau Commune is home to many standard hotels and resorts, attracting high-income domestic and international tourists. Meanwhile, Quan Lan Commune has many motels mainly built and operated by local people, so the scale and quality are not high, and will be suitable for ordinary tourists such as students.
c. Time of exploitation of Quan Lan Island Commune:
Quan Lan tourism is seasonal due to weather and climate conditions and festivals only take place on certain days of the year, specifically in spring. In Quan Lan commune, the period from April to June and from September to November is considered the best time to visit Quan Lan because the cultural tourism activities are mainly associated with festivals taking place during this time.
Minh Chau island commune:
Tourism exploitation time is all year round, because this is a place with a number of tourist attractions with diverse ecosystems such as Bai Tu Long National Park Research Center, Tram forest, Turtle Laying Beach, so besides coming to the beach for tourism and vacation in the summer, Minh Chau will attract research groups to come for tourism combined with research at other times of the year.
d. Sustainability
The sustainability of ecotourism sites in Quan Lan and Minh Chau communes depends on the sensitivity of the ecosystems to climate changes.
landscape. In general, these tourist destinations have a fairly high level of sustainability, because they are natural ecosystems, planned and protected. However, if a large number of tourists gather at certain times, it can exceed the carrying capacity and affect the sustainability of the environment (polluted beaches, damaged trees, animals moving away from their habitats, etc.), then the sustainability of the above ecosystems (natural ecosystems, human ecosystems) will also be affected and become less sustainable.
e. Location and accessibility
Both island communes have ports to take tourists to visit from Van Don wharf:
- Quan Lan – Van Don traffic route:
Phuc Thinh – Viet Anh high-speed boat and Quang Minh high-speed boat, depart at 8am and 2pm from Van Don to Quan Lan, and at 7am and 1pm from Quan Lan to Van Don. There are also wooden boats departing at 7am and 1pm.
- Van Don - Minh Chau traffic route:
Chung Huong high-speed train, Minh Chau train, morning 7:30 and afternoon 13:30 from Van Don to Minh Chau, morning 6:30 and afternoon 13:00 from Minh Chau to Van Don.
f. Infrastructure
Despite receiving investment attention, the issue of infrastructure and technical facilities for tourism on Quan Lan Island is still an issue that needs to be resolved because it has a direct impact on the implementation of ecotourism activities. The minimum conditions for serving tourists such as accommodation, electricity, water, communication, especially medical services, and security work need to be given top priority. Ecotourism spots in Minh Chau commune are assessed to have better infrastructure and technical facilities for tourism because there are quite complete and synchronous conditions for serving tourists, meeting many needs of domestic and foreign tourists.
3.2.1.4. Determine assessment levels and assessment scales
Corresponding to the levels of each criterion, the index is the score of those levels in the order of 4, 3, 2, 1 decreasing according to the standard of each level: very attractive (4), attractive (3), average (2), less attractive (1).
3.2.1.5. Determining the coefficients of the criteria
For the assessment of DLST in the two communes of Quan Lan and Minh Chau islands, the students added evaluation coefficients to show the importance of the criteria and indicators as follows:
Coefficient 3 with criteria: Attractiveness, Exploitation time. These are the 2 most important criteria for attracting tourists to tourism in general and eco-tourism in particular, so they have the highest coefficient.
Coefficient 2 with criteria: Capacity, Infrastructure, Location and accessibility . Because the assessment area is an island commune of Van Don district, the above criteria are selected by the author with appropriate coefficients at the average level.
Coefficient 1 with criteria: Sustainability. Quan Lan has natural and human-made ecotourism sites, with high biodiversity and little impact from local human factors. Most of the ecotourism sites are still wild, so they are highly sustainable.
3.2.1.6. Results of DLST assessment on Quan Lan island
a. Assessment of the potential for natural tourism development
For Minh Chau commune:
+ Natural tourism attractiveness is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined as average (2 points) and the coefficient is quite important (coefficient 2), then the score of Capacity criterion is 2 x 2 = 4.
+ Exploitation time is long (4 points), the most important coefficient (coefficient 3) so the score of the Exploitation time criterion is 4 x 3 = 12.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is assessed as good (3 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 3 x 2 = 6 points.
The total score for evaluating DLST in Minh Chau commune according to 6 evaluation criteria is determined as: 12 + 4 + 12 + 4 + 4 + 6 = 42 points
Similar assessment for Quan Lan commune, we have the following table:
Table 3.3: Assessment of the potential for natural ecotourism development in Quan Lan and Minh Chau communes
Attractiveness of self-tourismof course
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
CommuneMinh Chau
12
12
4
8
12
12
4
4
4
8
6
8
42/52
Quan CommuneLan
6
12
6
8
9
12
4
4
4
8
4
8
33/52
b. Assessment of the potential for humanistic tourism development
For Quan Lan commune:
+ The attractiveness of human tourism is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined to be large (3 points) and the coefficient is quite important (coefficient 2), then the score of the Capacity criterion is 3 x 2 = 6.
+ Mining time is average (3 points), the most important coefficient (coefficient 3) so the score of the Mining time criterion is 3 x 3 = 9.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points.
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is rated as average (2 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 2 x 2 = 4 points.
The total score for evaluating DLST in Quan Lan commune according to 6 evaluation criteria is determined as: 12 + 6 + 6 + 4 + 4 + 4 = 36 points.
Similar assessment with Minh Chau commune we have the following table:
Table 3.4: Assessment of the potential for developing humanistic eco-tourism in Quan Lan and Minh Chau communes
Attractiveness of human tourismliterature
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Quan CommuneLan
12
12
6
8
9
12
4
4
4
8
4
8
39/52
Minh CommuneChau
6
12
4
8
12
12
4
4
4
8
6
8
36/52
Basically, both Minh Chau and Quan Lan localities have quite favorable conditions for developing ecotourism. However, Quan Lan commune has more advantages to develop ecotourism in a humanistic direction, because this is an area with many famous historical relics such as Quan Lan Communal House, Quan Lan Pagoda, Temple worshiping the hero Tran Khanh Du, ... along with local festivals held annually such as the wind praying ceremony (March 15), Quan Lan festival (June 10-19); due to its location near the port and long exploitation time, the beaches in Quan Lan commune (especially Quan Lan beach) are no longer hygienic and clean to ensure the needs of tourists coming to relax and swim; this is also an area with many beautiful landscapes such as Got Beo wind pass, Ong Phong head, Voi Voi cave, but the ability to access these places is still very limited (dirt hill road, lots of gravel and rocks), especially during rainy and windy times; In addition, other natural resources such as mangrove forests and sea worms have not been really exploited for tourism purposes and ecotourism development. On the contrary, Minh Chau commune has more advantages in developing ecotourism in the direction of natural tourism, this is an area with diverse ecosystems such as at Rua De Beach, Bai Tu Long National Park Conservation Center...; Minh Chau beach is highly appreciated for its natural beauty and cleanliness, ranked in the top ten most beautiful beaches in Vietnam; Minh Chau commune is also home to Tram forest with a large area and a purity of up to 90%, suitable for building bridges through the forest (a very effective type of natural ecotourism currently applied by many countries) for tourists to sightsee, as well as for the purpose of studying and researching.
Figure 3.1: Thenmala Forest Bridge (India) Source: https://www.thenmalaecotourism.com/(August 21, 2019)
3.2.2. Using SWOT matrix to evaluate Quan Lan island tourism
General assessment of current tourism activities of Quan Lan island is shown through the following SWOT matrix:
Table 3.5: SWOT matrix evaluating tourism activities on Quan Lan island
Internal agent
Strengths- There is a lot of potential for tourism development, especially natural ecotourism and humanistic ecotourism.- The unskilled labor force is relatively abundant.- resource environmentunpolluted, still
Weaknesses- Poorly developed infrastructure, especially traffic routes to tourist destinations on the island.- The team of professional staff is still weak.- Tourism products in general
quite wild, originalintact
general and DLST in particularalone is monotonous.
External agents
Opportunity- Tourism is a key industry in the socio-economic development strategy of the province and Van Don economic zone.- Quan Lan was selected as a pilot area for eco-tourism development within the framework of the green growth project between Quang Ninh province and the Japanese organization JICA.- The flow of tourists and especially ecotourism in the world tends toincreasing
Challenge- Weather and climate change abnormally.- Competition in tourism products is increasingly fierce, especially with other localities in the province such as Ha Long, Mong Cai...- Awareness of tourists, especially domestic tourists, about ecotourism and nature conservation is not high.
Through summary analysis using SWOT matrix we see that:
To exploit strengths and take advantage of opportunities, it is necessary to:
- Diversify products and service types (build more tourism routes aimed at specific needs of tourists: experiential tourism immersed in nature, spiritual cultural tourism...)
- Effective exploitation of resources and differentiated products (natural resources and human resources)
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s3 { color: #0D0D0D; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -3pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -2pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -1pt; } div.maincontent .s9 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s11 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s12 { color: black; font-family:Symbol, serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s13 { color: black; font-family:Wingdings; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s14 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s15 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s16 { color: black; font-family:Cambria, serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s17 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s18 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s19 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s20 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 10pt; } div.maincontent .s21 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 11pt; } div.maincontent .s22 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s23 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s24 { color: #212121; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; tex
Identify Rating Levels and Rating Scales
zt2i3t4l5ee
zt2a3gstourism,quan lan,quang ninh,ecology,ecotourism,minh chau,van don,geography,geographical basis,tourism development,science
zt2a3ge
zc2o3n4t5e6n7ts
of the islanders. Therefore, this indicator will be divided into two sub-indicators:
a1. Natural tourism attractiveness a2. Cultural tourism attractiveness
b. Tourist capacity
The two island communes in Quan Lan have different capacities to receive tourists. Minh Chau Commune is home to many standard hotels and resorts, attracting high-income domestic and international tourists. Meanwhile, Quan Lan Commune has many motels mainly built and operated by local people, so the scale and quality are not high, and will be suitable for ordinary tourists such as students.
c. Time of exploitation of Quan Lan Island Commune:
Quan Lan tourism is seasonal due to weather and climate conditions and festivals only take place on certain days of the year, specifically in spring. In Quan Lan commune, the period from April to June and from September to November is considered the best time to visit Quan Lan because the cultural tourism activities are mainly associated with festivals taking place during this time.
Minh Chau island commune:
Tourism exploitation time is all year round, because this is a place with a number of tourist attractions with diverse ecosystems such as Bai Tu Long National Park Research Center, Tram forest, Turtle Laying Beach, so besides coming to the beach for tourism and vacation in the summer, Minh Chau will attract research groups to come for tourism combined with research at other times of the year.
d. Sustainability
The sustainability of ecotourism sites in Quan Lan and Minh Chau communes depends on the sensitivity of the ecosystems to climate changes.
landscape. In general, these tourist destinations have a fairly high level of sustainability, because they are natural ecosystems, planned and protected. However, if a large number of tourists gather at certain times, it can exceed the carrying capacity and affect the sustainability of the environment (polluted beaches, damaged trees, animals moving away from their habitats, etc.), then the sustainability of the above ecosystems (natural ecosystems, human ecosystems) will also be affected and become less sustainable.
e. Location and accessibility
Both island communes have ports to take tourists to visit from Van Don wharf:
- Quan Lan – Van Don traffic route:
Phuc Thinh – Viet Anh high-speed boat and Quang Minh high-speed boat, depart at 8am and 2pm from Van Don to Quan Lan, and at 7am and 1pm from Quan Lan to Van Don. There are also wooden boats departing at 7am and 1pm.
- Van Don - Minh Chau traffic route:
Chung Huong high-speed train, Minh Chau train, morning 7:30 and afternoon 13:30 from Van Don to Minh Chau, morning 6:30 and afternoon 13:00 from Minh Chau to Van Don.
f. Infrastructure
Despite receiving investment attention, the issue of infrastructure and technical facilities for tourism on Quan Lan Island is still an issue that needs to be resolved because it has a direct impact on the implementation of ecotourism activities. The minimum conditions for serving tourists such as accommodation, electricity, water, communication, especially medical services, and security work need to be given top priority. Ecotourism spots in Minh Chau commune are assessed to have better infrastructure and technical facilities for tourism because there are quite complete and synchronous conditions for serving tourists, meeting many needs of domestic and foreign tourists.
3.2.1.4. Determine assessment levels and assessment scales
Corresponding to the levels of each criterion, the index is the score of those levels in the order of 4, 3, 2, 1 decreasing according to the standard of each level: very attractive (4), attractive (3), average (2), less attractive (1).
3.2.1.5. Determining the coefficients of the criteria
For the assessment of DLST in the two communes of Quan Lan and Minh Chau islands, the students added evaluation coefficients to show the importance of the criteria and indicators as follows:
Coefficient 3 with criteria: Attractiveness, Exploitation time. These are the 2 most important criteria for attracting tourists to tourism in general and eco-tourism in particular, so they have the highest coefficient.
Coefficient 2 with criteria: Capacity, Infrastructure, Location and accessibility . Because the assessment area is an island commune of Van Don district, the above criteria are selected by the author with appropriate coefficients at the average level.
Coefficient 1 with criteria: Sustainability. Quan Lan has natural and human-made ecotourism sites, with high biodiversity and little impact from local human factors. Most of the ecotourism sites are still wild, so they are highly sustainable.
3.2.1.6. Results of DLST assessment on Quan Lan island
a. Assessment of the potential for natural tourism development
For Minh Chau commune:
+ Natural tourism attractiveness is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined as average (2 points) and the coefficient is quite important (coefficient 2), then the score of Capacity criterion is 2 x 2 = 4.
+ Exploitation time is long (4 points), the most important coefficient (coefficient 3) so the score of the Exploitation time criterion is 4 x 3 = 12.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is assessed as good (3 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 3 x 2 = 6 points.
The total score for evaluating DLST in Minh Chau commune according to 6 evaluation criteria is determined as: 12 + 4 + 12 + 4 + 4 + 6 = 42 points
Similar assessment for Quan Lan commune, we have the following table:
Table 3.3: Assessment of the potential for natural ecotourism development in Quan Lan and Minh Chau communes
Attractiveness of self-tourismof course
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
CommuneMinh Chau
12
12
4
8
12
12
4
4
4
8
6
8
42/52
Quan CommuneLan
6
12
6
8
9
12
4
4
4
8
4
8
33/52
b. Assessment of the potential for humanistic tourism development
For Quan Lan commune:
+ The attractiveness of human tourism is determined to be very attractive (4 points) and the most important coefficient (coefficient 3), so the score of the Attractiveness criterion is 4 x 3 = 12.
+ Capacity is determined to be large (3 points) and the coefficient is quite important (coefficient 2), then the score of the Capacity criterion is 3 x 2 = 6.
+ Mining time is average (3 points), the most important coefficient (coefficient 3) so the score of the Mining time criterion is 3 x 3 = 9.
+ Sustainability is determined as sustainable (4 points), the important coefficient is the average coefficient (coefficient 1), so the score of the Sustainability criterion is 4 x 1 = 4 points.
+ Location and accessibility are determined to be quite favorable (2 points), the coefficient is quite important (coefficient 2), the criterion score is 2 x 2 = 4 points.
+ Infrastructure is rated as average (2 points), the coefficient is quite important (coefficient 2), then the score of the Infrastructure criterion is 2 x 2 = 4 points.
The total score for evaluating DLST in Quan Lan commune according to 6 evaluation criteria is determined as: 12 + 6 + 6 + 4 + 4 + 4 = 36 points.
Similar assessment with Minh Chau commune we have the following table:
Table 3.4: Assessment of the potential for developing humanistic eco-tourism in Quan Lan and Minh Chau communes
Attractiveness of human tourismliterature
Capacity
Mining time
Sustainability
Location and accessibility
Infrastructure
Result
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Point
DarkMulti
Quan CommuneLan
12
12
6
8
9
12
4
4
4
8
4
8
39/52
Minh CommuneChau
6
12
4
8
12
12
4
4
4
8
6
8
36/52
Basically, both Minh Chau and Quan Lan localities have quite favorable conditions for developing ecotourism. However, Quan Lan commune has more advantages to develop ecotourism in a humanistic direction, because this is an area with many famous historical relics such as Quan Lan Communal House, Quan Lan Pagoda, Temple worshiping the hero Tran Khanh Du, ... along with local festivals held annually such as the wind praying ceremony (March 15), Quan Lan festival (June 10-19); due to its location near the port and long exploitation time, the beaches in Quan Lan commune (especially Quan Lan beach) are no longer hygienic and clean to ensure the needs of tourists coming to relax and swim; this is also an area with many beautiful landscapes such as Got Beo wind pass, Ong Phong head, Voi Voi cave, but the ability to access these places is still very limited (dirt hill road, lots of gravel and rocks), especially during rainy and windy times; In addition, other natural resources such as mangrove forests and sea worms have not been really exploited for tourism purposes and ecotourism development. On the contrary, Minh Chau commune has more advantages in developing ecotourism in the direction of natural tourism, this is an area with diverse ecosystems such as at Rua De Beach, Bai Tu Long National Park Conservation Center...; Minh Chau beach is highly appreciated for its natural beauty and cleanliness, ranked in the top ten most beautiful beaches in Vietnam; Minh Chau commune is also home to Tram forest with a large area and a purity of up to 90%, suitable for building bridges through the forest (a very effective type of natural ecotourism currently applied by many countries) for tourists to sightsee, as well as for the purpose of studying and researching.
Figure 3.1: Thenmala Forest Bridge (India) Source: https://www.thenmalaecotourism.com/(August 21, 2019)
3.2.2. Using SWOT matrix to evaluate Quan Lan island tourism
General assessment of current tourism activities of Quan Lan island is shown through the following SWOT matrix:
Table 3.5: SWOT matrix evaluating tourism activities on Quan Lan island
Internal agent
Strengths- There is a lot of potential for tourism development, especially natural ecotourism and humanistic ecotourism.- The unskilled labor force is relatively abundant.- resource environmentunpolluted, still
Weaknesses- Poorly developed infrastructure, especially traffic routes to tourist destinations on the island.- The team of professional staff is still weak.- Tourism products in general
quite wild, originalintact
general and DLST in particularalone is monotonous.
External agents
Opportunity- Tourism is a key industry in the socio-economic development strategy of the province and Van Don economic zone.- Quan Lan was selected as a pilot area for eco-tourism development within the framework of the green growth project between Quang Ninh province and the Japanese organization JICA.- The flow of tourists and especially ecotourism in the world tends toincreasing
Challenge- Weather and climate change abnormally.- Competition in tourism products is increasingly fierce, especially with other localities in the province such as Ha Long, Mong Cai...- Awareness of tourists, especially domestic tourists, about ecotourism and nature conservation is not high.
Through summary analysis using SWOT matrix we see that:
To exploit strengths and take advantage of opportunities, it is necessary to:
- Diversify products and service types (build more tourism routes aimed at specific needs of tourists: experiential tourism immersed in nature, spiritual cultural tourism...)
- Effective exploitation of resources and differentiated products (natural resources and human resources)
div.maincontent .p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent p { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; margin:0pt; } div.maincontent .s1 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s2 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 13pt; } div.maincontent .s3 { color: #0D0D0D; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s4 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s5 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s6 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -3pt; } div.maincontent .s7 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -2pt; } div.maincontent .s8 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; vertical-align: -1pt; } div.maincontent .s9 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s10 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s11 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s12 { color: black; font-family:Symbol, serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s13 { color: black; font-family:Wingdings; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s14 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s15 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 9pt; vertical-align: 5pt; } div.maincontent .s16 { color: black; font-family:Cambria, serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s17 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 14pt; } div.maincontent .s18 { color: #080808; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s19 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s20 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 10pt; } div.maincontent .s21 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: bold; text-decoration: none; font-size: 11pt; } div.maincontent .s22 { color: black; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; text-decoration: none; font-size: 11pt; } div.maincontent .s23 { color: black; font-family:"Times New Roman", serif; font-style: italic; font-weight: normal; text-decoration: none; font-size: 14pt; } div.maincontent .s24 { color: #212121; font-family:"Times New Roman", serif; font-style: normal; font-weight: normal; tex -
 Expert Survey Results on Research Scales
Expert Survey Results on Research Scales -
 Research Projects of Regional and World Countries
Research Projects of Regional and World Countries -
 Discussion of Quantitative Research Results on the Impact of Control Variables on Financial Performance
Discussion of Quantitative Research Results on the Impact of Control Variables on Financial Performance -
 The Research Results of the Thesis Are Consistent with the Stated Scientific Hypothesis, and the Tasks of the Topic Have Been Solved.
The Research Results of the Thesis Are Consistent with the Stated Scientific Hypothesis, and the Tasks of the Topic Have Been Solved.
Citri
(Source: Cubero and Graham, 2002)
1.3.4.2. Introduction to disease-causing genes
Gene pth A
pth A is a member of the avrBs3 / pth A gene family. This gene is responsible for the virulence of X. axonopodis pv . citri on citrus (Brunings and Gabriel, 2003). However, when transferred to other Xanthomonas bacteria , it lost its virulence. The pth A gene carries multiple identical repeat sequences of 102 bp in the central region (Yang and Gabriel, 1995). The pth A gene is the trigger for the characteristic symptoms of citrus canker, including excessive cell proliferation and necrotic lesions (Duan et al., 1999).
In non-host plants such as tobacco, legumes and cotton, the pth A gene causes a hypersensitive response characterized by cell necrosis of infected tissues. Citrus canker is not induced when tested for expression of the avrb6 gene , which has approximately 97% sequence homology to the pth A gene. The avrb6 gene was cloned from X. campestris pv. malvacearum, which causes cotton leaf blight (De Feyter et al., 1993).
pth A, pth B, pth C and pth W are homologs of each other, they were isolated from X. axonopodis pv. citri strain A, X. axonopodis pv. aurantifolii strains B and C, X. axonopodis pv. citri strain Aw , respectively. All of these genes determine the pathogenicity of the bacteria on citrus trees. Of which, gene pth A limits the host range. The two genes pth C and pth W do not limit the host range (Brunings and Gabriel, 2003).
Gene hrp W
The interaction between parasites and their hosts depends on the protein system. This is especially true for the interaction of Gram-negative plant pathogens with their hosts. The hrp (Hypersensitive Response and Pathogenicity) gene cluster encodes proteins that constitute the type III secretion system (T3S-Type III secretion). This type III secretion system plays a role in distributing toxic effector proteins into the host cell (Leach and White, 1996; Dunger et al., 2005). Then, if the host is susceptible, a disease phenotype will appear, for resistant plants, a hypersensitive response will occur. The hrp gene cluster is part of a pathogenic island located on the main chromosome, spanning more than 23 kb and containing a region with a G + C content different from the rest of the bacterial genome (Brunings and Gabriel, 2003; Hacker and Kaper, 2000).
The hrp gene also carries mobile genetic elements.
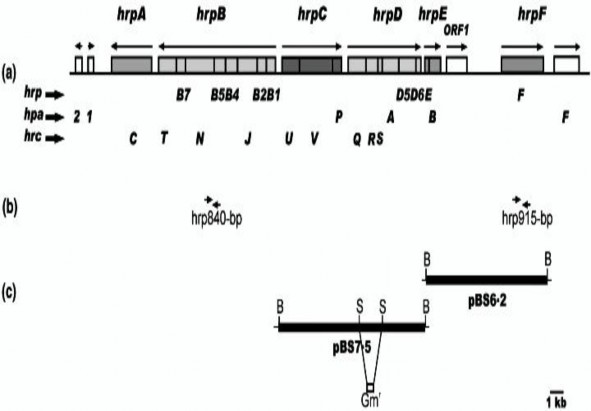
Figure 1.8. The hrp gene cluster of X. axonopodis pv . citri
(Source: Dunger et al., 2005)
On the bacteria X. axonopodis pv. citri, people have identified the following types of hrp genes: hrp G, hrp A, hrp B, hrc V, hrp B1, hrp D6, hrp B2, hrc U, hrc W, hrp B4, hrc N. In which, the genes belonging to this hrp gene cluster are located in clusters except for hrp W (Alegria et al., 2004). Based on this characteristic, people have designed specialized primers to quickly and accurately identify and detect the pathogenic strain using PCR technique. The model of the hrp gene cluster organization based on the genome sequence of the bacteria X. axonopodis pv. citri 306 is shown in Figure 1.8.
1.3.5. Some research results in the world and in the country on the bacteria Xanthomonas axonopodis pv. citri and canker disease caused by the bacteria Xanthomonas axonopodis pv. citri on citrus trees
Studies on biochemical characteristics
Bacterial canker disease on citrus is one of the serious diseases, affecting the quality and commercial value of the fruit and is very difficult to manage when it spreads.
spread rapidly. The disease has been present in most citrus growing countries in the world (Sharma and Sharma, 2009) and is currently one of the quarantine objects in trade. Therefore, the isolation and identification of the pathogen have been studied and reported in many countries around the world. The causative agent of canker disease on lemon trees has been reported to be X. axonopodis pv. citri and X. axonopodis pv. aurantifolia depending on the geographical region and host (Gottwald et al., 2002) . According to previous studies, on Nutrient agar (NA) medium, bacterial colonies are creamy yellow or pale yellow, convex, and slimy (Sujata and Sai, 2010; Ismail et al., 2014; Kharde et al., 2018; Isokar et al., 2020; Mahmood et al., 2020). On Nutrien glucose agar (NGA) medium, bacterial colonies are yellow, slimy, and convex (Al-saleh et al., 2014; Manyam and Nargund, 2020). On Yeast extract Dextrose (YDC) agar medium, bacterial colonies are yellow, round, slimy, and convex (Arshiya et al., 2014; Mahmood et al., 2020). On Galactose yeast peptone (GYP) medium, bacterial colonies are yellow, small, slimy, and shiny (Shehzadi and Nazi, 2019). On Yeast extract peptone glucose agar medium, bacterial colonies are pale yellow to yellow, medium in size, convex, shiny, and slimy (Haider et al., 2020). In general, previous studies have shown that Nutrient agar (NA) medium is often used to isolate the causative agent of canker disease on lemon trees. However, on NA medium, bacteria of the genus Pantoea also have a yellow color similar to the genus Xanthomonas (Mbega et al., 2012). Therefore, isolation and identification of colony characteristics are only a preliminary screening step. It is necessary to have a re-isolation step after assessing the pathogenicity to compare with the initial colony characteristics.
Studies evaluating the pathogenicity of Xanthomonas axonopodis pv. citri
After isolating the pathogen, assessing the pathogenicity of the agent is a mandatory procedure in determining the pathogen on plants. The time of disease manifestation depends on the bacterial strain and the host plant. X. axonopodis pv. punicae bacteria showed 100% leaf ulcer symptoms at 21 days after inoculation (Chand and Kishun, 1991). After inoculation on leaves, the initial lesions caused by X. axonopodis pv. citri bacteria showed the following symptoms: round, slightly raised, small, light green spots. Later, the inoculated lesions turned dark brown, solid, with a yellow halo around them (Haider et al., 2020). According to Abhang et al. (2015), Kharde et al. (2018)
and Haider et al. (2020), the highly virulent X. axonopodis pv. citri ( Xac ) 1 bacteria created wet areas and developed full symptoms of canker disease when inoculated on lemon leaves at 10 to 15 days after inoculation. When inoculated on citrus leaves, X. axonopodis pv. citri bacteria showed canker disease at 6 to 11 days after inoculation. At 6 days after inoculation, the lesion size reached 1.8 - 2.5 mm and continued to increase in size up to 20 days after inoculation (Arshiya et al., 2014; Mustafa et al., 2015). However, in the report of Katkar et al. (2016), in 15 strains of X. axonopodis pv. citri strains were inoculated on lemon leaves, the strains Xac -III, Xac -V, Xac -VII, Xac -XI, Xac -XIII and Xac -XIV were highly virulent, showing typical disease symptoms such as the formation of white crystalline callus at 7 to 9 days after inoculation. The strains Xac -I, Xac -II and Xac -IV were less virulent, showing disease symptoms at 13 to 16 days after inoculation. However, in the report of Mahmood et al. (2020), the symptoms of leaf canker after inoculation were observed quite a long time after 3 weeks after inoculation. Meanwhile, in the report of Manyam and Nargund (2020), 5 strains of X. axonopodis pv . citri had a fairly early disease manifestation time of 5 to 8 days after inoculation. The disease symptoms were quite typical: in the early stage, the lesions appeared on the leaf surface, with a yellow halo. Then the lesion increases in size (5-10 mm), the lesion cracks and becomes dark brown.
In general, studies evaluating pathogenicity use the method of causing wounds on leaves. Then, spray the prepared bacterial solution with a concentration of 10 6 - 10 8 cfu/mL on the leaves (with wounds) and incubate. The results of the disease manifestation in the early stage: the diseased area forms a wet area, with a small yellow halo around it. Then, the diseased area increases in size, the diseased area becomes hard, hard, cracked and turns dark brown, surrounded by a large yellow halo. The time of disease appearance is about 5 to 16 days after inoculation. In which, strains with strong virulence have an early disease appearance time from 5 to 9 days after inoculation. In addition, some studies also use the leaf cutting method. However, the wounding method gives better evaluation results, reported in most studies. After evaluating pathogenicity, the diseased areas are re-isolated on the original environment. The re-isolation results will be compared with the original isolation results.
Research results on identification of Xanthomonas axonopodis bacteria based on biochemical characteristics
To identify the pathogen, in addition to isolating and identifying the shape of the colony, identification based on biochemical characteristics is one of the scientific bases for species identification. When observed under a microscope, X. axonopodis bacteria are rod-shaped, thin, Gram (-), measuring 1.5 ÷ 2.0 x 0.5 ÷ 0.75 µm, have unipolar flagella, and are mobile (Das, 2003; Graham et al., 2004; Sing and Thind, 2014). On Nutrient agar (NA) isolation medium, the colony morphology of the bacteria is characterized by: pale yellow color, small, slimy, and protruding colonies (Graham et al., 2004; Shehzadi and Nazi, 2019). Biochemical characteristics of X. axonopodis bacteria . axonopodis : negative in oxidase, indole test, positive in catalase reactions, hydrolysis of gelatin, aesculin, hydrolysis of starch, casein, sucrose utilization, H 2 S production (Sujata and Sai, 2010; Islam et al., 2014; Arshiya et al., 2014; Haider et al., 2020, Isokar et al., 2020 ) , tolerant to 1, 2 and 3% salt and incapable of utilizing nitrate, citrate (Bharadwaj et al., 2014; Shehzadi and Nazi, 2019; Mahmood et al., 2020). In addition, X. axonopodis is capable of growing at 36 o C, not growing at 40 o C, hydrolyzing tween 20, 80, using L-arabinose, L-rahamnose, sucrose (Al-Saleh et al., 2014), producing fluorescent pigments (Mubeen et al., 2015), producing acid and gas, using Dextrose as carbon source (Kharde et al., 2018).
In general, previous studies have shown that X. axonopodis pv. citri bacteria have the following biochemical characteristics: Gram (-), rod-shaped, size from 1.5 ÷ 2.0 x 0.5 ÷ 0.75 µm, has unipolar flagella, obligate aerobic growth, negative for oxidase tests, fluorescent pigment production, arginine hydrolysis, urea, indole test, nitrate production, methyl red, voges-proskauer; positive for casein, starch, gelatin, tween 20, 80 hydrolysis tests, uses L-arabinose, L-rahamnose, sucrose and tolerates 1, 2 and 3% salt.
Studies on identification of Xanthomonas axonopodis bacteria causing ulcers based on sequences of specific gene regions
According to morphological classification based on colony shape, color, biochemical characteristics, species in the genus Xanthomonas can be identified. However, this method takes a lot of time and is prone to confusion. PCR techniques, DNA sequencing and
Other molecular characteristics are widely used as a standard tool for the detection, identification and classification of species in the genus Xanthomonas for rapid and accurate results (Del Campo et al., 2009).
Cubero and Graham (2002) suggested that based on the rDNA sequence region with primer pair 2/3, it is enough to identify two groups of X. axonopodis species : group 1 ( X. axonopodis pv. citri ) causing type A ulcer and group 2 ( X. axonopodis pv. aurantifolii ) causing type B ulcer and
C. Based on the pth A sequence region, X. axonopodis pv. citri bacteria can be detected quickly and accurately . The combination of two primer pairs 2/3 and Jpth1/Jpth2 can overcome the shortcomings when detecting the A W strain . The primer pair J-RXg/J-RXc2 is not suitable for detecting X. axonopodis pv. citri in plant samples. Sixty-four bacterial strains from lemon trees in Saudi Arabia were isolated and identified by 16S rDNA region sequencing with primer pairs 27R and 1492R. As a result, compared with the 16S rDNA region sequences on Genbank worldwide, the isolated bacterial strains were identified as X. citri subsp. citri (Al-Saleh et al., 2014). Also with the primer pair 27F/1492R to identify the 16S rDNA sequence region, the samples isolated on leaves and orange fruits had PCR band results with a size of 1500 bp, compared with the sequence on the world Genebank similar to X. citri subsp. citri (CP008989), the similarity level is 99% (Li et al., 2015). In addition, to quickly and accurately identify the bacteria X. axonopodis pv. citri, Park et al. (2006) suggested that the specific primer pair XacF/XacR could be used to identify the hrp W gene region. Manyam and Nargund (2020) used the primer pair Xc-lipF/Xc-lipR to identify the species of citrus canker isolates based on the est A gene region. The PCR results were 777 bp in size, and identified 5 isolates as X. axonopodis pv. citri.
In general, the Xanthomonas species causing canker disease on lemon trees can be identified based on the 16S rDNA region with the primer pair 27R and 1492R. In addition, the gene regions pth A, hrp W and est A were also used to identify the Xanthomonas species causing canker disease on lemon trees. According to Cubero and Graham (2002), the combination of sequencing the 16S rDNA region and the pth A gene sequence region with the primer pair Jpth1/Jpth2 quickly and accurately identified the species X. axonopodis pv. citri and overcame the shortcomings when identifying subspecies.
Bui et al. (2009) studied the distribution of X. citri pv. citri in Southeast Asia,
Among 577 X. citri pv. citri isolates from samples collected in 14 northern (Hanoi, Hung Yen, Nghe Han, Ha Tinh and Phu Tho) and southern (Can Tho, Long An, Dong Nai, Tien Giang, Vinh Long, Ben Tre, Dong Thap, Vung Tau and Lam Dong) provinces of Vietnam, 60 strains were from Mexican lemons. Intermediate-PCR (IS-LM) analysis using primers targeting three insertion sequences (1) was performed on all Vietnamese strains and on the reference strains X. citri pv. citri -A, -A* and X. citri pv. aurantifolii , IS-LM-PCR showed that all Vietnamese isolates were genotype A. Amplified fragment length polymorphism (AFLP) analysis was performed on 84 X. citri pv. citri , including 22 strains from Mexican limes, AFLP was performed using SacI/MspI and four primer pairs (MspI +1 [A, C, T, G] and 5′-SacI + C) for selective amplification). The results showed that all X. citri pv. citri strains in Vietnam were genetically related to the bacterial strain A.
In summary, canker disease caused by Xanthomonas sp. bacteria on lemon trees is quite common in many countries around the world and in all lemon growing areas in the country, causing serious damage, affecting the commercial quality of the fruit. In the world, many studies have reported that the bacteria causing canker disease on lemon trees are identified as X. axonopodis based on morphological, biochemical, disease manifestation characteristics, sequencing of the 16S rDNA region, pth A region and hrp W region using specialized primer pairs. However, domestic studies on the causative agent of canker disease are still limited. Currently, only the study of Bui et al. (2009) has been recorded, the group of authors reported that the bacteria causing canker disease on Mexican lemon trees in 14 northern and southern provinces is X. citri pv. citri .
1.4. Overview of Euphorbia tirucalli L.
1.4.1. Plant classification
Euphorbia tirucalli L. ( E. tirucalli L.) belongs to the genus Euphorbia, one of the
8,000 species in the family Euphorbiaceae. This is a small, shrub endemic to the tropics with pencil-like branches. Therefore, E. tirucalli is also known as the pencil tree or some other names such as: blue coral tree, fishbone tree, and giao tree.





