2002; Schuurman et al., 2004).
Check the quality of DNA after extraction ( Schuurman et al., 2004).
Amplification of target gene region by PCR reaction : For bacterial strains belonging to the genus Bacillus sp., PCR reaction was performed and identified with the common ITS primer pair for the 16S-RNA region 27F (5' AGATTTGATCCTGGCTCAG 3') and 1492R (5' GGTTACCTTGTTACGACTT 3') (Schuurman et al., 2004).
PCR product testing: PCR products are tested by electrophoresis on 1.5% agarose gel in 0.5 X TBE buffer at 100 V for 35 minutes. Then stained with Gel Red, the results are viewed under UV light and qualified PCR samples will be sequenced, processed by fastPCR and compared with sequences at NCBI/BLAST (Schuurman et al., 2004).
Maybe you are interested!
-
 Isolation and Identification of Bacteria (Frerichs and Millar (1983, 1993))
Isolation and Identification of Bacteria (Frerichs and Millar (1983, 1993)) -
 Study on nitrogen-metabolizing bacteria in the bottom of the lobster farming area Panulirus sp. for aquaculture - 2
Study on nitrogen-metabolizing bacteria in the bottom of the lobster farming area Panulirus sp. for aquaculture - 2 -
 Internal Combustion Engine Vibration Isolation System
Internal Combustion Engine Vibration Isolation System -
 Chemical study and identification of some groups of substances in the red-top tree (cratoxylum prunifolium kurtz) - 14
Chemical study and identification of some groups of substances in the red-top tree (cratoxylum prunifolium kurtz) - 14 -
 Chemical study and identification of some groups of substances in the red-top tree (cratoxylum prunifolium kurtz) - 12
Chemical study and identification of some groups of substances in the red-top tree (cratoxylum prunifolium kurtz) - 12
Sequencing the amplified gene segment: Send the PCR product to Nam Khoa Company (address 793/58 Tran Xuan Soan, Tan Hung Ward, District 7, Ho Chi Minh City). Sequencing is performed using the Sanger method, 1977. Compare the obtained sequence with the gene bank to determine the name of the bacterial species surveyed.
Genealogical tree construction: Collect reference sequences available on GenBank along with Assessment Number as the basis for constructing the genealogical tree. Use the Alignment tool of MEGA X software to align the sequences using the ClustalW method, the Alignment result is used as the basis for constructing the genealogical tree. Proceed to construct the genealogical tree using MEGA X software using the Neighbor Joining method with a bootstrap coefficient of 1000 (Kurma et al., 2018)

2.4.2. Isolation and identification of ammonia-metabolizing bacteria (AOB)
2.4.2.1. Medium for isolating ammonia-metabolizing bacteria (Appendix 1.3.1)
Bacterial isolation medium: ammonium-calcium-carbonate medium was used to isolate bacteria based on the method of Ehrlich (1975).
Bacterial enrichment culture medium was prepared according to the medium formula of Lewis and Pramer (1958) and MacDonad and Spokes (1980).
Griess – Ilosway reagent (Nguyen Duc Luong et al., 2006)
2.4.2.2. Steps to isolate ammonia-metabolizing bacteria
Test for the presence of ammonia-metabolizing bacteria: Sludge samples were
Stir well and transfer 10g of mixed mud into a bottle containing 90mL of sterile distilled water, shake well and then transfer the bacterial suspension into the ammonium liquid medium.
– calcium carbonate with NaCl added at concentrations of 1.5%, 2%, 2.5%, 3% and 3.5%, diluted to many concentrations. Falcol tubes containing this bacterial suspension solution and negative control tubes (no bacterial solution added) were incubated in the dark at 28 o C on a shaker for about 21 days. Every 3 days, the presence of ammonia-metabolizing bacteria in the sludge sample was checked using Griess – Ilosway reagent to check the conversion of NH 4 + to NO 2 - (Pham Thi Tuyet Ngan and Nguyen Huu Hiep, 2010).
Positive determination for ammonia-metabolizing bacteria: Reagent solutions A, B, C are mixed in a 1:1:1 ratio and added to 5 mL of bacterial suspension, shaken well and left for 5 minutes to see the results. Positive tubes have a dark or light pink color, negative control tubes do not give a color reaction (Pham Thi Tuyet Ngan and Nguyen Huu Hiep, 2010).
Method for determining AOB bacterial density according to MPN: The number of positive or negative times of the sludge sample is recorded and the Mac Crady statistical table is consulted to deduce the estimated value of the number of microorganisms in the sample. The unit of bacterial density is MPN/g (Ehrlich, 1975).
Enrichment and isolation of bacteria: To obtain high bacterial density and eliminate mixed bacteria, 20 mL of enrichment medium containing NaCl was added to falcon tubes. Then 1 mL of bacterial suspension solution determined to have the presence of AOB group was transferred to falcon tubes containing enrichment medium. The tubes were incubated in the dark at 28 0 C on a shaker for 21 days. After 7 days, 0.53 mg (NH 4 ) 2 SO 4 solution and 0.005 mL 1M NaOH were added to the tubes of the above medium, and the presence of NO 2 - was checked in the tubes of the bacterial isolation medium using Griess – Ilosway reagent. After 3 days, the possibility of heterotrophic bacterial contamination in those tubes was checked using TSA bacterial isolation medium (supplemented with NaCl). If no heterotrophic bacteria grow on TSA medium, it proves that pure ammonia-metabolizing bacteria have been obtained. Pipet 100 µl of bacterial suspension into a plate containing ammonium-calcium-agar medium.
carbonate was incubated for 2 - 4 days to isolate AOB bacteria (Lewis and Pramer, 1958; MacDonad and Spokes, 1980, Pham Thi Tuyet Ngan and Nguyen Huu Hiep, 2010).
Bacterial culture storage : After the AOB bacteria group is purified, the bacterial culture will be kept in a slant tube on LB agar medium, or stored at -80°C with 15% glycerol (Montras et al., 2008), and preserved in a liquid medium containing 0.53 mg (NH4)2SO4 and 0.005 mL 1M NaOH (Aleem and Alexander, 1960; Pham Thi Tuyet Ngan, 2012).
2.4.2.3. Biochemical identification method of AOB bacteria
Gram staining method and biochemical reaction: according to Sharmin and Rahman, 2007.
Qualitative determination of ammonia metabolism ability of AOB bacteria strains: Add 1 mL of isolated bacterial suspension solution into ammonium - calcium - carbonate medium, incubate and shake at 28 0 C for 10 days. After 10 days, test the ability to convert NH 4 + to NO 2 - of AOB bacteria group using NH 4 /NH 3 Sera test kit.
Method of using API 20E and API 20NE kits: After determining the ability to metabolize ammonia, the bacterial strains are subjected to biochemical reactions according to the instructions of Biomerieux, France (The global health network, 2013). According to the manufacturer's instructions, the API 20E kit is used for gram-negative bacteria with negative oxidase reactions, the API 20NE kit is used for gram-negative bacteria with positive oxidase reactions.
2.4.2.4. Molecular biological identification method of AOB bacteria
Extract DNA and check DNA quality after extraction : perform each step as in the method of Bacillus sp.bacteria
Amplify the target gene region by PCR reaction : For AOB bacterial strains, perform PCR reaction and identify with the common primer pair of 16S region - rRNA 27F (5' AGATTTGATCCTGGCTCAG 3') and 515 R (5'-TACCGCGGC
TGC TGG CAC-3') (Schuurman et al., 2004).
Check PCR products: perform each step as in section 2.4.1.3.
Sequencing the amplified gene segment: perform similarly to section 2.4.1.3.
Build genetic tree: do the same as section 2.4.1.3.
2.4.3. Method for isolating nitrite-metabolizing bacteria (NOB)
2.4.3.1. Environment for isolating nitrite-metabolizing bacteria
Nitrite – calcium – carbonate medium for isolation of nitrite-metabolizing bacteria based on the method of Ehrlich, 1975.
The enrichment medium for nitrite-metabolizing bacteria was prepared according to the medium of Aleem and Alexandre (1960).
2.4.3.2. Steps for isolating nitrite-metabolizing bacteria (NOB)
Check for the presence of nitrite-metabolizing bacteria: perform the steps as in section 2.4.2.2 on nitrite-calcium-carbonate medium supplemented with NaCl at different concentrations. Falcol tubes containing this bacterial suspension solution and negative control tubes (no bacterial solution added) are incubated in the dark at 28 0 C on a shaker for about 21 days. After incubation, check for the presence of NO 2 - using Griess-Ilosway reagent (Pham Thi Tuyet Ngan, 2012).
Positive determination for nitrite-metabolizing bacteria: Perform the steps as in section 2.4.2.2, the reaction between the bacterial suspension solution and the Griess-Ilosway reagent is white, indicating the presence of nitrite-metabolizing bacteria (NOB) in the environment, while the negative control tubes are dark pink (Pham Thi Tuyet Ngan, 2012).
Method for determining NOB bacterial density: similar to section 2.4.2.2
Enrichment and bacterial isolation: perform the same as section 2.4.2.2 on nitrate - calcium - carbonate medium, 20 µM NO 2 - solution and 0.02 mL sterilized 20% KOH are added to the above culture medium tubes every 4 days (Aleem and Alexander, 1960).
Store the bacterial culture as described in section 2.4.2.2 and also store in liquid medium containing solutions containing 20 µM NO 2 - and 0.02 mL 20% KOH (Aleem and Alexander, 1960; Pham Thi Tuyet Ngan, 2012).
2.4.3.3. Biochemical identification method of NOB bacteria
Gram staining method and biochemical reaction : perform similarly
2.4.2.3
Qualitative determination of nitrite metabolism ability of NO OB bacterial strains : Bacterial strains were cultured and grown in nitrite - calcium - carbonate medium, similar to section 2.4.2.3. The bacterial culture after growth was evaluated for nitrite treatment ability using Sera's NO 2 test kit.
2.4.3.4. Molecular biological identification method of NOB bacteria
Extract DNA and check DNA quality after extraction : perform as in section 2.4.2.4
Amplify the target gene region by PCR reaction : use the same primer pair according to section 2.4.2.4
PCR product testing: perform the same as section 2.4.2.4.
Sequencing the amplified gene segment: perform similarly to section 2.4.2.4.
Building a genetic tree: do the same as section 2.4.2.4.
2.4.4. Survey on the metabolism of ammonium, nitrite, and nitrate of selected bacterial strains
-
-
+ -
The selected bacterial strains were grown in LB medium until reaching a density of 10 7 CFU/mL, then added to ammonia quantification medium containing NH 4 10 mg/l, nitrite quantification medium containing NO 2 0.53 mg/l, Nitrate Broth medium with NO 3 content of 13.34 mg/l, at room temperature. Samples were collected at 12 to 48 hours ( Bacillus sp. bacteria group ), samples were collected at 1 day, 2 days, 3 days, 4 days, 5 days, 6 days, 7 days (AOB group), from 12 hours to 72 hours (NO 3 metabolism survey ) (Phatthongkleang, 2019; Sang, 2020).
+
-
-
NH 4 content in samples was determined by Phenat method. NO 2 content in samples was determined by Napthylamine method. NO 3 content was determined by Salicylate method (APHA, 2012) (Appendix 1.5). Quantitative results were converted to treatment efficiency, from which bacterial strains with high ammonia, nitrite, nitrate metabolism ability were selected to create biological products. Quantitative results will be converted to treatment efficiency according to the following formula:
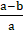
H = x 100%
In which: H is the treatment efficiency (%); a is the initial substance content (mg/l); b is the substance content after the i-th time (mg/l).
2.4.5. Survey on salt tolerance of bacterial strains
The salt tolerance of bacterial strains with ammonia, nitrite, and nitrate metabolism properties was investigated by: culturing bacterial strains at a density of 1.2 x 10 8 CFU/mL (OD 600nm = 1) in LB medium for each group of bacteria, supplemented with different concentrations of NaCl salt 2%, 3%, 4%, 5%, 6%, 7% ( Phatthongkleang, 2019) at room temperature for 48 hours. After culturing, collect the bacterial solution and measure OD with a spectrophotometer at a wavelength of 600 nm. The control is a salt-supplemented medium without bacteria.
2.5. Content 2: Creating liquid and powdered microbial preparations
2.5.1. Survey of factors affecting the culture and optimization of fermentation medium composition to create liquid preparations of bacterial strains
2.5.1.1. Survey of factors affecting the biomass growth of three bacterial strains.
Investigation of the influence of growth medium on bacterial growth
Inoculate bacterial colonies from the seed tube into a test tube containing autoclaved distilled water (121 o C/15 minutes), shake well for 10-20 minutes, measure OD at 600 nm wavelength to use the linear correlation line to deduce the bacterial cell density (log.CFU/mL). Then, add 1% of the bacteria with a density of 107 CFU /mL to 3 types of growth media: TSB, NB, LB (media composition in the appendix). After 24 hours of culture at 37 o C, count the colony density and calculate the bacterial density to select the optimal growth media for each bacterial strain.
The monitoring index for each experiment examining the influencing factors is the bacterial density (CFU/mL) by indirect colony counting method on agar plates and using the Log base 10 function in Microsoft Excel software according to the formula lga = b where a is the microbial density (CFU/mL), b is the conversion factor to 10.lg for statistical processing by 1-factor ANOVA analysis method through MSTATC software (Statistical Aspects of Microbiological Criteria Related to Foods, 2016).
Investigation of the effect of seed density on bacterial biomass growth
After selecting the growth medium, continue to base on the linear correlation line of microbial density and OD 600nm to prepare bacterial densities of 10 6 , 10 7 , 10 8 , 10 9 CFU/mL, then add 1% seed ratio with different bacterial densities into the tested growth medium, shake for 24 hours, temperature 37 o C, count colonies in each treatment to select the highest output bacterial density.
Investigation of the effect of time on bacterial biomass growth
Conduct the experiment with the environment and seed density surveyed in the above experiment, changing the growth time to 24 hours, 36 hours, 48 hours, 60 hours, 72 hours. Count the colonies after each time point to choose the most optimal growth time.
2.5.1.2. Survey of factors affecting the process of creating liquid preparations. Survey of the effect of seed loading ratio on the process of bacterial biomass multiplication on liquid production medium.
Bacteria after proliferation under conditions such as environment, seed density,
The time examined in section 2.5.1.1 and the density adjusted to reach 10 8 CFU/mL will be
inoculated into a conical flask containing 500 mL of production medium (appendix), with seed loading ratios of 1.0%, 2.5%, 3.5%, and 5.0%, respectively, shaking at 150 rpm at 30 o C for 24 hours. Calculate the bacterial density, thereby finding the best seed ratio.
Investigation of the effect of culture time on bacterial biomass multiplication on liquid production medium.
Conduct experiments with the determined inoculation rate, survey the following time points: 12 hours, 24 hours, 36 hours and 48 hours, evaluate the influence of culture time on the bacterial biomass multiplication process in the production environment, and select the culture time for the next experiment.
Investigation of the effect of temperature on the process of bacterial biomass multiplication in liquid production medium .
Conduct the experiment with the determined inoculum loading rate and time, shaking the bacterial production medium at 150 rpm at temperatures of 30 o C, 33 o C, 35 o C, 37 o C. Determine the bacterial density at each temperature level to choose the best culture temperature for the next experiment.
Investigation of the effect of pH on bacterial biomass multiplication in liquid production medium.
With the seed loading rate, time, and temperature determined in the above experiment, shake the bacterial production medium at 150 rpm at pH 6; 6.5; 7; 7.5. From there, determine the bacterial density at each pH level to choose the best culture pH for the experiment.
Investigation and screening of nitrogen and carbon sources affecting bacterial biomass multiplication process in liquid production medium.
Liquid TSB production medium is based on TSB growth medium with some changes in mineral composition and nitrogen and carbon content. With the initial carbon source in TSB production medium being glucose with a content of 10g/L, the carbon sources are changed to maltodextrin and molasses, sucrose with 10g/L.
After selecting the best carbon source, continue to select the nitrogen source. In the TSB production environment, the initial nitrogen source is peptone 10 g/L, changing the nitrogen sources to yeast extract, (NH 4 ) 2 SO 4 and NaNO 3 with 10 g/L.
After proliferation and density adjustment to reach 10 8 CFU/mL, bacteria will be inoculated into each medium to select carbon and nitrogen sources with optimal seed loading ratio, temperature, pH and culture time determined from previous experiments. Density count
The bacteria then select the carbon and nitrogen sources that mainly influence the process.
bacterial biomass
2.5.1.3. Optimization of the biomass media composition of three bacterial strains using response surface methodology (RMS).
Screening of significant factors by Plackett – Burman design of Bacillus licheniformis B85 and Pseudomonas stutzeri KL15
When conducting a single factor study, the two bacterial strains Bacillus licheniformis B85 and Pseudomonas stutzeri KL15 both used molasses and yeast extract as the best carbon and nitrogen sources. Plackett-Burman design was used to screen the main parameters in a large number of factors of the optimization process and allowed the evaluation of the factors, each factor was tested at two levels: low (-1) and high (+1). The factors selected for this study were molasses, yeast extract, K 2 HPO 4 , MgSO 4 , NaCl and CaCl 2 listed in Table 2.1.





