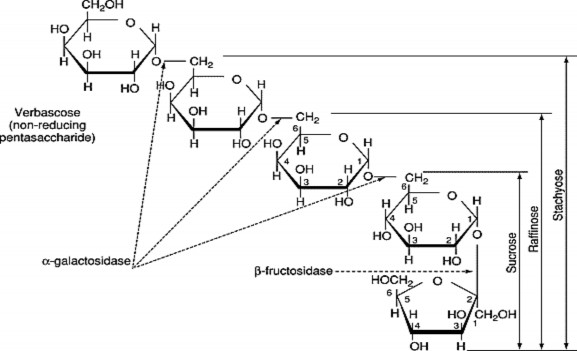
Figure 1.3 . Structure and relationships of sugars in the Raffinose family
O -α- d -Galactopyranosyl-(1→ 6) - [ O -α- d -galactopyranosyl- (1→ 6)] 2 - O -α- d -glucopyranosyl- (1→ 2) β- d -frutofuranoside. The arrows point out
sites hydrolyzed by enzymes [12] .
Maybe you are interested!
-
 Structure of Tay Folk Song Texts Considered According to Successive Relationships
Structure of Tay Folk Song Texts Considered According to Successive Relationships -
 Impact of Financial Market Development on Enterprise Capital Structure by National Institutional Quality
Impact of Financial Market Development on Enterprise Capital Structure by National Institutional Quality -
 The impact of financial market development on the capital structure of listed enterprises in the ASEAN Economic Community - 28
The impact of financial market development on the capital structure of listed enterprises in the ASEAN Economic Community - 28 -
 The impact of financial structure on the performance of commercial banks - 1
The impact of financial structure on the performance of commercial banks - 1 -
 Comparison of Geographical Conditions, Structure of Culture and Tourism Activities
Comparison of Geographical Conditions, Structure of Culture and Tourism Activities
RFOs are common storage substances in soybean seeds, which affect the moisture retention capacity of the seeds and the stress tolerance of the plant [13]. Some recent studies have shown that soybean seeds of cultivated varieties contain high levels of stachyose and raffinose [14]. In addition, some studies on soybean seeds with low levels of stachyose and raffinose or total raffinose (RFOs) compared to wild plants, did not lead to significant differences in stress tolerance or seed germination [15]. In purebred soybean varieties, the main component of RFOs is stachyose (30-60 g/1 kg), followed by raffinose (2-9 g/1 kg) [16]. Kumar's study also showed that soybean seeds contain 6.4-25.3 mM/kg raffinose and 20.9-71.0 mmol/kg stachyose [17]. In mature seeds, RFOs are mainly in the form of stachyose, these compounds are derived from sucrose and have a positive effect on energy metabolism. However, this is a group of sugars that are difficult to digest for humans and monogastric animals [10].
Recent studies have shown that RFOs are an energy source that affects seed germination, and inhibition of RFOs metabolism significantly reduces seed germination ability [18]. The stachyose content in seeds is directly proportional to the ability to withstand
The moisture content of soybean seeds, for young seeds that are dehydrated, will slow down the accumulation of RFOs [19]. These indigestible oligosaccharides pass through the digestive system from the stomach to the small intestine, then they are fermented by anaerobic microorganisms residing in the large intestine. The fermentation process produces CO 2 and CH 4 gases that disrupt digestive activity leading to flatulence and diarrhea in monogastric animals [20].
Interest in the factors causing this change stems from the agricultural significance of RFOs. Due to their poor metabolism in the digestive systems of humans and monogastric animals, the removal of RFOs from crop seeds is still under active research [21].
Raffinose
Raffinose (RFO) is a trisaccharide composed of galactose, glucose and fructose. It can be found in beans, cabbage, Brussels sprouts, broccoli, asparagus, other vegetables and whole grains. RFO has a higher molecular weight than Stachyose, Verbascose and Ajugose Raffinose, which can be hydrolyzed to D- galactose and sucrose by the enzyme α-galactosidase (α-GAL), an enzyme not present in the human digestive tract [22]. Scientists have suggested that RFO is considered as a protective agent for seed embryos during moisture production and storage in a dry state [13], however there is no evidence of a relationship between their accumulation and desiccation tolerance. RFO is often accumulated in vegetative tissues as a plant antibody to help plants resist biotic and abiotic stresses [23].
Stachyose
One of the most abundant tetrasaccharides in plants is stachyose. Stachyose consists of four sugar units: two α-D-galactose units, one α-D-glucose unit and one β-D-fructose unit linked as follows gal(α1→6) gal (α1→6) glc (α1↔2β) fru. Stachyose was first isolated from the rhizome of Stachys tuberifera , coexisting with RFO and other related oligosaccharides in various organs of many plant species. In woody plants, cucurbits and legumes, this sugar exists in vacuoles, roots, seeds... with storage and transportation functions in the plant.
Intermediate cells are the major sites of RFO and stachyose synthesis.
In Catalpa and Buddleia leaves , stachyose is synthesized from the combination of RFO and galactinol through photosynthesis. The enzyme raffinose synthase (RS) cannot synthesize stachyose, a specific enzyme exists in plant sources that synthesizes this tetrasaccharide. This enzyme was isolated from several sources and was found to catalyze the following reaction [24]:
[ 14 C] Galactinol + raffinose →[ 14 C] stachyose + myo-inositol
However, the displacement reaction is freely reversible, but stachyose synthesis is preferred [25]:
Stachyose + [ 14 C] raffinose ⇌ [ 14 C] stachyose + raffinose
In plants, stachyose is synthesized during leaf development and synthesis reaches its peak in mature leaves. Stachyose is transported to all other parts of the plant via the phloem, except for other mature leaves. Stachyose synthesis occurs even in old and partially yellowed leaves.
In seeds, stachyose is metabolized during germination. In winter hardy plants, stachyose and related oligosaccharides provide rigidity; their content varies seasonally with the size of their cell walls. Stachyose is mainly used as a sweetener or as an oligosaccharide. Stachyose is not completely digestible by humans.
Verbascose
Verbascose is a pentasaccharide present in the phloem and storage organs of many plants, especially in legume seeds [26]. This group of sugars is very low in seeds, so studies on the physiological function of verbascose in seeds are limited. Verbascose may act as a protective substance during drought-stressed seed maturation or as a carbon reserve for seed germination [27]. The content of verbascose in seeds can vary widely, specifically studies on the variation in verbascose content in faba beans ( Vicia faba ) ranged from 1.1% to 9.9% of seed dry matter [28]. In a survey of bean ( Pisum sativum ) genotypes, verbascose values ranged from undetectable to 3.1% of dry matter.
Early studies showed that verbascose biosynthesis proceeds by the transfer of a galactose unit from galactinol [O‐α‐d‐galactopyranosyl‐(1→1))l‐myo‐inositol] to stachyose. The isolation of the enzyme verbascose synthase
(VBS) from pea seeds has been demonstrated by scientists to be a multifunctional stachyose synthase [29]. The enzyme operates on a ping-pong mechanism. In the first half of the reaction, a galactose unit is transferred from galactinol to the enzyme, forming an intermediate enzyme-galactose complex:
Galactinol + E ⇔ myo ‐ inositol + [E ‐ Gal] (1)
In the second half of the reaction, the galactose unit is transferred to raffinose (STS activity) or stachyose (VBS activity):
[E ‐ Gal] + raffinose ⇔ E + stachyose (2) [E ‐ Gal] + stachyose ⇔ E + verbascose (3)
All reactions are reversible, occurring at a single active site. Therefore, the reversal of the half-reactions (Equation 2) and half-reactions (Equation 3) can be combined as a complete catalytic cycle, producing verbascose and raffinose from stachyose as the sole substrate:
Stachyose + stachyose ⇔ raffinose + verbascose (4)
In a study on the accumulation of RFOs in two soybean cultivars, it was shown that the cultivar also influences the accumulation of verbascose. Specifically, the accumulation of RFOs was studied in the mature seeds of two bean ( Pisum sativum ) lines, the seeds of line SD1 accumulated stachyose as dominant while verbascose appeared to be absent. However, the seeds of line RRRbRb had some amount of verbascose accumulated together with stachyose. The increase in verbascose in developing RRRbRb seeds was associated with increased verbascose synthase activity and galactinol-dependent mRNAs encoding the enzymes [30].
1.2.2 RFOs biosynthesis pathway in soybean and involved enzymes
1.2.2.1 Biosynthetic pathway of RFOs in soybean
The biosynthesis of RFOs in plants involves a series of reactions that are carried out to transfer the galactosyl functional group (active group) from functional group donors (including: UDPD-galactose, galactinol and from the RFOs compounds themselves) to typical functional group acceptors such as sucrose, raffinose or compounds with high isotopes to it (Figure 1.4) [31]. Many studies have shown that the synthesis of RFOs usually takes place in leaves, specifically the mesophyll layer of the cell membrane and intermediate cells located in the phloem layer, and then will be transported to other parts.
in plants and accumulates most in seeds [30]. The synthesis of RFOs in plants occurs as follows:
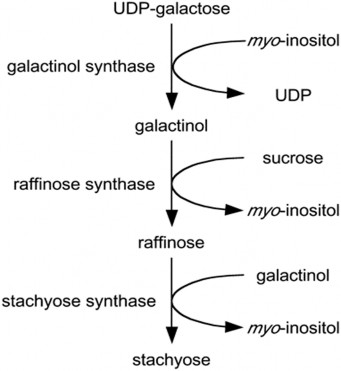
Step one (1): under the influence of the GOLS enzyme, galactinol is formed from two initial components, UDP-galactose and myo-inositiol.
Step two (2): the enzyme raffinose synthase (RS) catalyzes the transfer of the galactosyl group from galactinol to the carbon 6 (C6) position of the glucose unit in the sucrose molecule, creating an α-1,6-galactosidic bond to form the trisaccharide compound raffinose [32].
Step three (3) of RFOs biosynthesis: the enzyme stachyose synthase catalyzes the transfer of a galactosyl unit from galactinol to the 6-carbon position of the galactose unit in the raffinose component to form a compound with four stachyose monosaccharide units [33].
In soybean, raffinose biosynthesis occurs in the developing seed and is catalyzed by raffinose synthase 2 (RS2), encoded by Glyma06g18890 [14]. RS2 catalyzes the reaction between sucrose and galactinol to form raffinose and myo-inositol. The subsequent conversion of raffinose to stachyose and stachyose to verbascose produces a complex of raffinose oligosaccharides, however the content
(1)
(2)
Verbascose in soybean seeds is negligible [17].
(3)
Figure 1.4. Biosynthetic pathway of RFOs [33].
1.2.2.2 Enzymes involved in the biosynthetic pathway
Raffinose synthase (RS)
RS is the second enzyme involved in the synthesis of RFOs from galactinol and sucrose, first purified from Vicia faba seeds , followed by wheat germ and soybean seeds [33]. RS has a neutral pH optimum, catalyzing the galactosylation, or inversion of sucrose from galactinol to produce raffinose and myo -inositol [33].
Stachyose synthase (STS)
This is the third enzyme in the RFOs synthesis pathway. The enzyme is involved in the synthesis of stachyose from galactinol and raffinose and has been proposed as a regulatory enzyme for RFOs synthesis, based on the findings that STS activity in leaves increases in parallel with increased fruit set in cucumber and watermelon; STS was first isolated and characterized from the seeds of Phaseo lus vulgaris and subsequently purified and extracted from Vicia faba seeds and from the leaves of several cucurbits and Ajuga . The enzyme is optimally active at neutral pH [35].
Verbascose Synthase
This enzyme catalyzes the formation of DP, higher RFOs, which has received little attention. Although the photosynthetic formation of [14C]-verbascose from 14CO, has been observed in the leaves of Lamium maculatum and Xerosicyos danguyi , no verbascose synthesis was found in in vitro leaf extracts [33].
1.2.3 Gene encoding for Galactinol synthase ( GOLS) in soybean
GOLS is an important enzyme in the biosynthesis of raffinose by catalyzing the formation of galactinol from myo-inositol and UDP-Galactose. Loss of function of the gene encoding GOLS reduces galactinol, stachyose and increases sucrose content has been studied in Arabidopsis . In addition, increasing the expression of genes encoding enzymes involved in galactinol biosynthesis ( AtGolS1, AtGolS2, AtGolS4) in Arabidopsis increased galactinol, raffinose concentrations and increased oxidative resistance [36]. However, studies on the function of this group of genes in soybean are still very limited. Mutation studies of each gene separately or together will be the basis for analyzing the function of
The genes encoding GOLS in soybean are also oriented for application in improving and enhancing the quality of soybean seeds, resulting in a reduction in the content of raffinose and starchyose in soybean seeds.
1.2.4 Study on changes in RFOs content in soybean plants
GOLS is the first enzyme involved in the biosynthesis of RFOs. GOLS catalyzes the conversion of L-myo-inositiol to galactinol, the first reaction in the biosynthesis of raffinose [31]. Overexpression of the galactinol synthase gene in transgenic Arabidopsis plants increased the endogenous levels of galactinol and raffinose under normal conditions [37].
RS catalyzes the conversion of sucrose into raffinose and subsequent high molecular weight compounds such as Stachyose, Verbascose and Ajugose. In soybean, three genes encoding RS have been identified, of which the gene encoding RS2 ( Glyma06g18890 ) has the strongest expression during soybean seed development [14]. Therefore, reducing the expression of this gene will increase the sucrose content while reducing the content of simple sugars such as raffinose and stachyose. Some soybean lines carrying the RS2 mutant gene have been confirmed to contain low levels of raffinose in seeds [38].
In addition, studies on soybean lines created by RNAi technology to reduce the expression of the RS2 gene also recorded a decrease in raffinose sugar in soybean seeds and an increase in the efficiency of digestion and energy absorption of poultry using this seed [15]. This shows an opportunity and important significance in studying the biosynthesis of raffinose in improving the quality of soybean seeds. In addition, STS is an enzyme that plays an important role in the biosynthesis of stachyose. Natural mutant lines showed that inhibition of this enzyme activity also increased the sugar content in the seeds and significantly reduced the stachyose content in the seeds.
[39].
1.3 Genome editing via CRISRP/Cas9 system
In 2011-2012, two independent research groups led by Dr. Emmanuelle Charpentier at Umea University and Dr. Jennifer Doudna at the University of California, Berkeley, jointly discovered the mechanism of action of the Cas enzyme and CRISPR. The CRISPR/Cas system was identified as an immune system in prokaryotes that is capable of fighting foreign genetic elements such as plasmid invasion and bacterial phages; CRISPR/Cas is also known as the genetic immune system.
The mechanism of action of Cas and CRISPR enzymes is based on RNA pairing with the spacer sequence on newly invaded DNA and helping Cas protein (CRISPR-associated) recognize and cut the DNA strand; in addition, the complex consisting of guide RNA and Cas protein can also cut foreign invading RNAs [41].
In January 2013, Doudna and Charpentier made a breakthrough in cutting a DNA sample from a human cell and replacing it with a different genetic code [42]. At the same time, another group of researchers from Harvard University and the Broad Institute independently announced that they had successfully developed a similar method [43]. The main features of the CRISPR/Cas system are: (1) creating a double-stranded break in DNA and (2) this break is precisely located by a guide crRNA segment. Scientists have likened CRISPR/Cas to “molecular scissors” that can cut out unwanted gene segments and replace them with new ones on demand [44]. In 2015, Science magazine voted CRISPR/Cas9 as the most important technology, starting a new era of biotechnology, helping to edit the genetic information of all cells quickly and accurately [45]. And two scientists, Doudna and Charpentier, with their work on CRISPR/Cas9, won the Nobel Prize in 2020.
Since then, the CRISPR/Cas system has been researched and applied by scientists in the fields of medicine, pharmaceuticals, agriculture, etc. This system is being applied in cancer treatment research by eliminating parts of the genome directly responsible for tumor growth, destroying mosquitoes carrying the Zika virus that causes yellow fever, treating blindness, and destroying HIV [46]. In addition, CRISPR/Cas is also used in research on genetic diseases through editing the genomes of human embryos; developing better drugs; improving the properties of agricultural plants or animals in livestock farming, etc. [47].
1.3.1. Operating principle of CRISRP/Cas system
This system is based on the “immune” mechanism of bacteria against foreign DNA molecules from viruses or plasmid DNA. Unlike the “immune” system based on restriction enzymes, the CRISPR/Cas system is based on RNA molecules to recognize and destroy foreign DNA. To protect bacteria from foreign DNA, bacteria insert a short piece of foreign DNA into their genomic DNA at the CRISPR repeat sequence region. This sequence region is transcribed and processed into RNA segments





