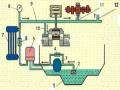
Figure 3.3. The phenomenon of insertion and deletion at the categorical position of A/H5N1 strains cl de 7 isolated in Vietnam
3.2.2. Sequencing the NA(N1) gene and analyzing the phylogenetic tree based on the nucleotide sequence of the N1 gene
The NA gene of avian influenza virus together with the HA gene creates the antigen on the surface of the capsid. Therefore, in parallel with the sequencing of the H gene, we also sequenced and analyzed the N1 gene for the 5 A/H5N1 clade influenza virus samples that we isolated. The names of the 0 virus strains for which we sequenced the N1 gene can be as follows:
- A/Chicken/Vietnam/NCVD-016/2008
- A/Chicken/Vietnam/NCVD-093/2008
- A/Chicken/Vietnam/NCVD-03/2008
- A/Chicken/Vietnam/NCVD-04/2008
- A/Chicken/Vietnam/NCVD-05/2008
The length of the N1 gene segment of the sequenced influenza A/H5N1clade 7 virus samples was 1346 nucleotides, encoding 448 amino acids, starting with the ATG codon. The nucleotide sequences of the N1 gene of these virus strains were compared with
The corresponding N1 gene sequences of some A/H5N1 influenza virus strains of Vietnam and China discovered in the period 1996-2010 include the A/goose/Guangdong 1 96 virus strain, which is considered the ancestor of the highly pathogenic A/H5N1 influenza virus. Along with the comparison of the differences in nucleotide sequences between the virus strains, we established a phylogenetic tree of the N1 gene based on the nucleotide sequences and the results are shown in Figure 3.4. The AH N1 virus strains used to establish the phylogenetic tree along with the AH N1 virus strains belonging to the Vietnamese clade are listed in Table 3.5.
Figure 3.4 shows that the results of phylogenetic analysis based on the N1 gene are also consistent with the results of phylogenetic comparison based on the H gene. That is, the N1 gene of the A/H5N1 clade 7 influenza virus strains isolated in Vietnam also has high similarity to the N1 gene of the influenza virus strains belonging to clade 7 in China and are grouped together on the phylogenetic tree.
Detailed observation of the family tree shows that among the five strains of A/H5N1 virus clade
7 Vietnamese isolates have 4 strains: A chicken Vietnam NCVD-03/2008, A/chicken/Vietnam/NCVD-04/2008, A/chicken/Vietnam/NCVD-05/2008 and A/Chicken/Vietnam/NCVD-016/2008 nmcm grouped together. Meanwhile, strain A/chicken/Vietnam/NCVD-093 is quite far away from this group and has a close similarity (nm close together on the phylogenetic tree) with the virus strain belonging to clade 7 of China isolated in 2003 (A/Beijing/01/2003).
The results of the comparison of the distribution of A/H5N1 influenza virus strains of clade 7 in the gene tree of H and N1 genes, it can be observed that there are also different evolutions of the N1 gene among the A/H5N1 influenza virus strains, which may be the A/chicken/Vietnam/NCVD-093 strain that is different from the remaining group of Vietnamese clade viruses. However, when compared with A/H5N1 influenza virus strains of other clades, it can be clearly seen that the A/H5N1 influenza viruses of clade 7 in China and Vietnam have high similarity and are completely different from other clades currently circulating in the region.
81
100
85
A/CHICKEN/VIETNAM/NCVD-05/2008 *
A/CHICKEN/VIETNAM/NCVD-04/2008 *
A/CHICKEN/VIETNAM/NCVD-03/2008 *
97
73
79
89 68
A/chicken/Shanxi/10/2006
A/CHICKEN/VIETNAM/NCVD-016/2008 *
A/chicken/Hebei/706/2005
A/chicken/Liaoning/A-1/2007 A/mallard/Huadong/hn/2005
A/chicken/Shanxi/2/2006
A/chicken/Ningxia/24/2006
20 amino acid fragment
Virus clade 7 HA
99 92
97
A/chicken/Hebei/126/2005 A/chicken/Hebei/102/2005
95
98 61
A/chicken/Hebei/326/2005 A/Beijing/January 2003 2003
A/CHICKEN/VIETNAM/NCVD-093/2008 *
A/Vietnam/1194/2004
46 A/Vietnam/1203/2004 A/chicken/Vietnam/8/2003
83
10050
91 70
A/chicken/Viet Nam/AG-010/2004 A/Vietnam/CL20/2004
A/chicken/Viet Nam/DT-015/2004 A/chicken/Bac Lieu/07-10/2007
100
95
A/Muscovy duck/Ca Mau/07-04/2007 A/duck/Viet Nam/38ST/2008
100
99
98
A/duck/Hue/V6/2010
A/duck/Viet Nam/TMU026/2009 A/chicken/Viet Nam/TMU025/2009
A/duck/Hai Phong /208/2006
71 A/chicken/Bac Giang/07-74/2007
100
55
A/duck/Ha Tinh/07-53/2007 A/chicken/Ha Nam/07-83/2007
100
51
A/duck/Vietnam/8/05 A/goose/Vietnam/3/05
Deletion of 19 amino acids
A/duck/Vietnam/1/2005
A/Hong Kong/485/1997
95
A/chicken/China/27402/1997
99
52 A/HongKong/486/97
72 A/HongKong/481/97
69 54
99
86
49
65
100
A/duck/Guangxi/2291/2004 A/duck/Guangxi/1378/2004 A/chicken/Viet Nam/Ncvd8/2003
A/Hong Kong/213/2003 A/HK/212/03 2003
40 A/Hong Kong/156/97
A/goose/Vietnam/113/2001
A/Pheasant/HongKong/FY155/01
99
A/goose/Hong Kong/437-6/1999
A/Goose/Guangdong/3/97
90 A/Chicken/Hong Kong/YU822.2/01
89 A/Goose/Guangdong/1/96
0.005
Figure 3.4. Phylogenetic tree of the N1 gene nucleotide sequence of A/H5N1 clade 7 strains (Established by neighbor-joining analysis using MEGA 5 software, boostrap value 1000)
irus is a virus /H5N1 clade 7 isolated from Vietnam
Table 3.5. Identification of A/H5N1 strains used in the establishment of the N1 genotype
Status
International trade | Species battery | Capacity set up | Place of isolation | S bank registration gene | |
1 | A/goose/Guangdong/1/96 | Goose | 1996 | China | AF144304 |
2 | A/chicken/Hongkong/YU822/01 | Chicken | 1997 | Hong Kong | AY221545 |
3 | A/Hongkong/156/97 | People | 1997 | Hong Kong | AF036357 |
4 | A/Hongkong/481/97 | People | 1997 | Hong Kong | AF102663 |
5 | A/Hongkong/485/1997 | People | 1997 | Hong Kong | AF102664 |
6 | A/Hongkong/486/1997 | People | 1997 | Hong Kong | AF102658 |
7 | A/chicken/China/27402/1997 | Chicken | 1997 | China | GU052122 |
8 | A/goose/Guangdong/3/97 | Goose | 1997 | China | AF364335 |
9 | A/goose/Hongkong/437-6/1999 | Goose | 1999 | Hong Kong | GU052021 |
10 | A/pheasan/Hongkong/FY115/01 | Hemorrhoids | 2001 | Hong Kong | AF509096 |
11 | A/goose/Vietnam/113/2001 | Goose | 2001 | Vietnam | EF541474 |
12 | A/Hongkong/212/2003 | People | 2003 | Hong Kong | AY575881 |
13 | A/Hongkong/213/2003 | People | 2003 | Hong Kong | AB557744 |
14 | A/Beijing/01/2003 | People | 2003 | China | EF587279 |
15 | A/chicken/Vietnam/8/2003 | Chicken | 2003 | Vietnam | DQ493042 |
16 | A/chicken/Vietnam/ncvd8/2003 | Chicken | 2003 | Vietnam | EF541468 |
17 | A/duck/Guangxi/1378/2004 | Duck | 2004 | China | DQ321016 |
18 | A/duck/Guangxi/2291/2004 | Duck | 2004 | China | DQ321022 |
19 | A/chicken/Vietnam/AG-010/2004 | Chicken | 2004 | Vietnam | DQ094278 |
20 | A/chicken/Vietnam/DT-015/2004 | Chicken | 2004 | Vietnam | DQ094279 |
21 | A/Vietnam/1194/2004 | People | 2004 | Vietnam | EF541466 |
22 | A/Vietnam/1203/2004 | People | 2004 | Vietnam | HM006761 |
23 | A/Vietnam/CL20/2004 | People | 2004 | Vietnam | DQ493071 |
24 | A/chicken/hebei/102/2005 | Chicken | 2005 | China | EU243134 |
25 | A/chicken/hebei/126/2005 | Chicken | 2005 | China | EU2431451 |
26 | A/chicken/hebei/326/2005 | Chicken | 2005 | China | DQ349118 |
27 | A/chicken/hebei/706/2005 | Chicken | 2005 | China | EU243127 |
28 | A/mallard/huadong/hn/2005 | Mallard | 2005 | China | EU195410 |
29 | A/goose/Vietnam/3/05 | Goose | 2005 | Vietnam | DQ366316 |
30 | A/duck/Vietnam/1/05 | Duck | 2005 | Vietnam | DQ366308 |
31 | A/duck/Vietnam/8/05 | Duck | 2005 | Vietnam | DQ366324 |
32 | A/chicken/ningxia/24/2006 | Chicken | 2006 | China | HM172205 |
33 | A/chicken/shanxi/10/2006 | Chicken | 2006 | China | HM172172 |
34 | A/chicken/shanxi/2/2006 | Chicken | 2006 | China | DQ914816 |
Maybe you are interested!
-
 Phenomenon that changes reaction rate - 2
Phenomenon that changes reaction rate - 2 -
 Damage Phenomenon, Causes, Remedies
Damage Phenomenon, Causes, Remedies -
 Phylogenetic Tree of Nucleotide Sequence of Gen M of CC Strain IA/h5N1 Clade 7 (Established by Neighbo-Joining Analysis Using Mega 5 Software, Gi T
Phylogenetic Tree of Nucleotide Sequence of Gen M of CC Strain IA/h5N1 Clade 7 (Established by Neighbo-Joining Analysis Using Mega 5 Software, Gi T -
 Esp Control Brake System Against Under- or Over-Revving Phenomenon
Esp Control Brake System Against Under- or Over-Revving Phenomenon -
 Phenomenon that changes reaction rate - 5
Phenomenon that changes reaction rate - 5

Status
International trade | Species battery | Year isolate | Place of isolation | S register gene bank | |
35 | A/duck/Haiphong/208/2006 | Duck | 2006 | Vietnam | GU052504 |
36 | A/chicken/Liaoning/a-1/2007 | Chicken | 2007 | China | HM172206 |
37 | A/chicken/Bacgiang/07-74/2007 | Chicken | 2007 | Vietnam | GU050469 |
38 | A/chicken/Baclieu/07-10/2007 | Chicken | 2007 | Vietnam | GU186763 |
39 | A/chicken/Hanam/07-83/2007 | Chicken | 2007 | Vietnam | GU050477 |
40 | A/muscovyduck/Camau/April 7, 2007 | Goose | 2007 | Vietnam | GU186747 |
41 | A/duck/Hatinh/07-53/2007 | Duck | 2007 | Vietnam | GU050430 |
42 | A/duck/Vietnam/38ST/2008 | Duck | 2008 | Vietnam | FJ812005 |
43 | A/chicken/Vietnam/TMU025/2009 | Chicken | 2009 | Vietnam | CY095702 |
44 | A/duck/Hue/V6/2010 | Duck | 2009 | Vietnam | JN935005 |
45 | A/duck/Vietnam/TMU026/2009 | Duck | 2009 | Vietnam | CY095705 |
The results of N1 gene sequencing showed that the A/H5N1clade 7 influenza virus strains in Vietnam had a deletion mutation, resulting in the loss of 20 amino acids at positions 49-68 in the stem region of the NA antigen molecule. This deletion mutation of 20 amino acids has also been observed in most influenza viruses detected and isolated in Vietnam and China from 2004 to present.
There is another mutation phenomenon on the NA antigen molecule, which is the deletion of 19 amino acids in the A/H5N1 influenza virus strains isolated from Hong Kong in 1999 (such as strain A Hongkong 169) and some A/H5N1 influenza virus strains isolated from Vietnam in 2000. However, until recently, the deletion mutation of 19 amino acids in A/H5N1 virus strains has not been seen anymore. Most of the classical highly pathogenic A/H N1 virus strains (such as strain A/goose/Guangdong/1/96) have the N1 gene with the maximum length (no amino acid deletion mutation) and this phenomenon has only been detected in A/H N1 viruses isolated until 2004.
The NA antigenic body plays an important role in the virulence of avian influenza viruses. It has been found that a 20-amino acid deletion mutation at position 49-68 is associated with high virulence (Zhou et al., 2006). This phenomenon was also found by Zhao et al., 2007 (Zhao et al., 2008) and that may represent the adaptation of influenza A/H5N1 virus to poultry (Atrosovich et al., 1999).
We also analyzed the genetic differences in the N1 gene at the nucleotide and amino acid levels of the AH N1 virus strains belonging to clade 7 collected in Vietnam and some A/H5N1 virus strains belonging to clade 7 in China (A/chicken/Shanxi/10/2006, A/chicken/Shanxi/2/2006, A/Beijing/01/2003) compared with some other AH N1 influenza virus strains from China (A/HongKong/156/97, A/Goose/Guangdong/1 96) and from Vietnam (A/VietNam/1194/2004, A/VietNam/1203/2004, A/goose/VietNam/113/2001, A/Goose/Guangdong/1/96). The results are presented in Table 3.6.
When comparing the level of genetic differences between the A/H5N1clade 7 influenza virus strains of Vietnam, we found that the differences between the four A/H5N1 clade 7 influenza viruses of Vietnam mentioned above, namely A/chicken/Vietnam/NCVD-03/2008, A/chicken/Vietnam/NCVD-04/2008, A/chicken/Vietnam/NCVD-05/2008 and A Chicken Vietnam NCVD-016 2008, were very low, ≤ 2% at the nucleotide level and ≤ 2.% at the amino acid level. As clearly seen in the family tree, this virus group has a relatively large difference compared to another virus strain of Vietnam that also belongs to the clade, A/chicken/Vietnam/NCVD-093/2008. The differences in nucleotide levels and amino acid levels between the group of 4 virus strains we mentioned above and the virus A/chicken/Vietnam/NCVD-093/2008 were 4. ±0.2% (4.5-4.9%) and
,6±0.3% ( ,3-6.0%).
The analysis results also showed that the A/chicken/Vietnam/NCVD-093 virus strain is most closely related to the A/Beijing/01/2003 virus strain, which was isolated from humans in 2003 in China. The differences are only 2.3% at the nucleotide level and 2.0% at the amino acid level.
The largest observable difference between the clade 7 influenza A/H5N1 viruses collected in Vietnam and the A/H N1 virus strains with a 19-amino acid deletion mutation (A/Hong_Kong/156/97 virus and A goose Vietnam 30 virus) was 12.3-1.1% nucleotide and 9.-10.4% amino acid levels.
Table 3.6. Genetic differences in the N1 gene of five A/H5N1 strains compared to a reference strain
Symbol
strain
Strain name | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | |
Amino acids (%) | |||||||||||||||
1 | A/Chicken/Vietnam/NCVD-016/2008 | 2.5 | 2.7 | 2.7 | 5.3 | 2.3 | 3.0 | 3.4 | 5.0 | 4.8 | 6.2 | 9.5 | 9.7 | 5.3 | |
2 | A/Chicken/Vietnam/NCVD-03/2008 | 1.9 | 0.7 | 0.4 | 5.5 | 2.0 | 2.5 | 3.6 | 5.8 | 5.5 | 6.2 | 9.7 | 9.9 | 5.3 | |
3 | A/Chicken/Vietnam/NCVD-04/2008 | 2.0 | 0.5 | 0.4 | 6.0 | 2.3 | 3.0 | 4.1 | 6.0 | 5.8 | 6.7 | 10.2 | 10.4 | 5.8 | |
4 | A/Chicken/Vietnam/NCVD-05/2008 | 2.0 | 0.4 | 0.4 | 5.5 | 2.3 | 2.7 | 3.6 | 6.0 | 5.8 | 6.2 | 10.0 | 10.2 | 5.3 | |
5 | A/Chicken/Vietnam/NCVD-093/2008 | 4.5 | 4.7 | 4.9 | 4.5 | 5.5 | 4.1 | 2.5 | 5.5 | 5.0 | 5.3 | 9.7 | 9.9 | 4.1 | |
6 | A/chicken/Shanxi/10/2006 | 1.6 | 1.4 | 1.6 | 1.7 | 4.5 | 2.7 | 3.6 | 5.5 | 5.5 | 6.5 | 10.2 | 10.4 | 5.3 | |
7 | A/chicken/Shanxi/2/2006 | 2.7 | 2.5 | 2.7 | 2.7 | 4.1 | 2.4 | 2.0 | 4.6 | 4.1 | 4.8 | 8.5 | 8.5 | 3.9 | |
8 | A/Beijing/01/2003 | 2.4 | 2.7 | 3.0 | 2.8 | 2.3 | 2.3 | 2.3 | 3.6 | 3.2 | 3.6 | 8.2 | 8.2 | 2.3 | |
9 | A/Viet_Nam/1194/2004 | 4.7 | 5.2 | 5.4 | 5.4 | 4.3 | 4.7 | 4.6 | 2.8 | 0.9 | 5.0 | 8.7 | 8.7 | 3.4 | |
10 | A/Viet_Nam/1203/2004 | 4.2 | 4.8 | 5.0 | 5.0 | 3.8 | 4.3 | 4.1 | 2.3 | 1.0 | 4.6 | 8.2 | 8.2 | 3.4 | |
11 | A/goose/Viet_Nam/113/2001 | 6.6 | 6.7 | 6.8 | 6.4 | 5.7 | 6.5 | 6.0 | 4.5 | 5.6 | 4.9 | 7.9 | 7.7 | 2.8 | |
12 | A/Hong_Kong/156/97 | 14.2 | 14.8 | 15.1 | 14.7 | 13.8 | 14.4 | 13.2 | 13.0 | 13.8 | 13.3 | 12.2 | 1.1 | 6.9 | |
13 | A/goose/Vietnam/3/05 | 13.0 | 13.4 | 13.7 | 13.3 | 12.3 | 13.0 | 11.9 | 11.4 | 12.3 | 11.7 | 10.9 | 1.8 | 6.9 | |
14 | A/Goose/Guangdong/1/96 | 5.2 | 5.6 | 5.9 | 5.5 | 4.3 | 5.3 | 5.0 | 3.0 | 3.6 | 3.3 | 3.2 | 11.1 | 9.8 | |
Nucleotides (%) | |||||||||||||||
77
The genetic differences between the influenza virus strains belonging to the clade we collected in Vietnam and the virus strain considered to be the ancestor of the highly pathogenic AH N1 influenza virus (A/Goose/Guangdong/1/96) were 4.3–.9% at the nucleotide level and 4.1–.8% at the amino acid level.
3.2.3. Sequencing gene M and analyzing the phylogenetic tree based on the nucleotide sequence of gene M
The genome of avian influenza virus type A in general and AH N1 virus consists of 8 segments, including 2 segments of HA and NA genes encoding 2 types of surface antigens that are significant for determining the subtype of the virus. At present, the highly pathogenic avian influenza virus strains AH N1 Southeast Asia have HA and NA genes originating and evolving from the precursor virus A Goose Guangdong 196 discovered in China. Meanwhile, the genes encoding the endogenous proteins (, PA, PB2 and PB1) come from many different sources. This has created recombination into many different genotypes (Guan et al., 2002; Li et al., 2004). Therefore, to further study the genetic characteristics of AH N1 virus strains belonging to the Vietnamese clade, we conducted gene sequencing of these virus samples.
The nucleotide sequence length of the sequenced gene is 983 bp (base pair) encoding for 2 gene segments 1 and 2. The sequence of the M gene of the virus strain we isolated was compared with some highly pathogenic AH N1 avian influenza virus strains detected in Asian countries, including the Guangdong 1 96 Avian Influenza virus strain, which is the ancestor of highly pathogenic avian influenza viruses and some previous Vietnamese virus strains. Based on the difference in the M gene sequence of the compared AH N1 virus strains, we established a phylogenetic tree to track and analyze the evolution of these virus strains compared with previous AH N1 influenza virus strains.
The list of AH N1 virus strains used for comparison and phylogenetic tree establishment is presented in Table 3.7.